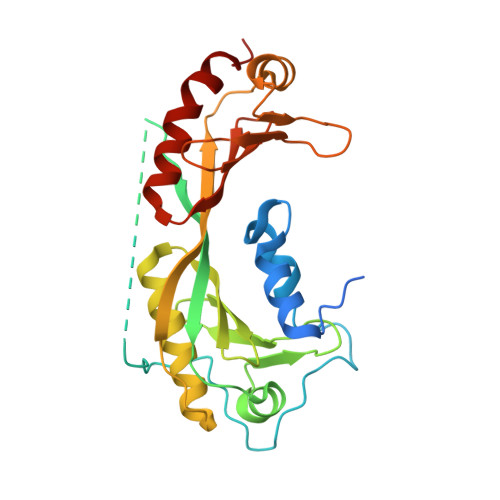
High-Resolution Structure of TBP with Taf1 Reveals Anchoring Patterns in Transcriptional Regulation
Anandapadamanaban, M., Andresen, C., Siponen, M., Kokubo, T., Ikura, M., Moche, M., Sunnerhagen, M.(2013) Nat Struct Mol Biol 20: 1008
- PubMed: 23851461 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2611
- Primary Citation Related Structures:
4B0A - PubMed Abstract:
The general transcription factor TFIID provides a regulatory platform for transcription initiation. Here we present the crystal structure (1.97 Å) and NMR analysis of yeast TAF1 N-terminal domains TAND1 and TAND2 bound to yeast TBP, together with mutational data. We find that yeast TAF1-TAND1, which in itself acts as a transcriptional activator, binds TBP's concave DNA-binding surface by presenting similar anchor residues to TBP as does Mot1 but from a distinct structural scaffold. Furthermore, we show how TAF1-TAND2 uses an aromatic and acidic anchoring pattern to bind a conserved TBP surface groove traversing the basic helix region, and we find highly similar TBP-binding motifs also presented by the structurally distinct TFIIA, Mot1 and Brf1 proteins. Our identification of these anchoring patterns, which can be easily disrupted or enhanced, provides insight into the competitive multiprotein TBP interplay critical to transcriptional regulation.
- Department of Physics, Chemistry and Biology, Linköping University, Linköping, Sweden.
Organizational Affiliation: