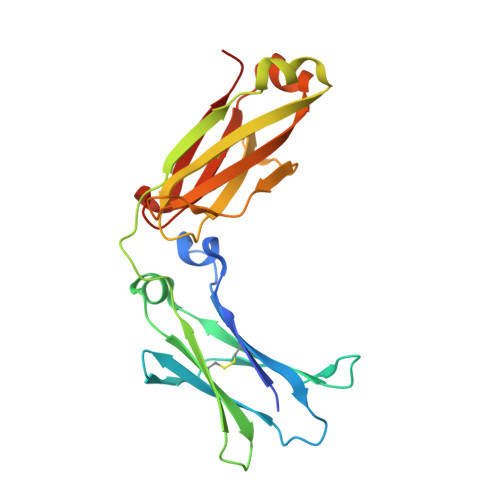
Structural Determinants of Unique Properties of Human Igg4-Fc
Davies, A.M., Rispens, T., Ooijevaar-Deheer, P., Gould, H.J., Jefferis, R., Aalberse, R.C., Sutton, B.J.(2014) J Mol Biology 426: 630
- PubMed: 24211234
- DOI: https://doi.org/10.1016/j.jmb.2013.10.039
- Primary Citation Related Structures:
4C54, 4C55 - PubMed Abstract:
Human IgG4, normally the least abundant of the four subclasses of IgG in serum, displays a number of unique biological properties. It can undergo heavy-chain exchange, also known as Fab-arm exchange, leading to the formation of monovalent but bispecific antibodies, and it interacts poorly with FcγRII and FcγRIII, and complement. These properties render IgG4 relatively "non-inflammatory" and have made it a suitable format for therapeutic monoclonal antibody production. However, IgG4 is also known to undergo Fc-mediated aggregation and has been implicated in auto-immune disease pathology. We report here the high-resolution crystal structures, at 1.9 and 2.35 Å, respectively, of human recombinant and serum-derived IgG4-Fc. These structures reveal conformational variability at the CH3-CH3 interface that may promote Fab-arm exchange, and a unique conformation for the FG loop in the CH2 domain that would explain the poor FcγRII, FcγRIII and C1q binding properties of IgG4 compared with IgG1 and -3. In contrast to other IgG subclasses, this unique conformation folds the FG loop away from the CH2 domain, precluding any interaction with the lower hinge region, which may further facilitate Fab-arm exchange by destabilisation of the hinge. The crystals of IgG4-Fc also display Fc-Fc packing contacts with very extensive interaction surfaces, involving both a consensus binding site in IgG-Fc at the CH2-CH3 interface and known hydrophobic aggregation motifs. These Fc-Fc interactions are compatible with intact IgG4 molecules and may provide a model for the formation of aggregates of IgG4 that can cause disease pathology in the absence of antigen.
- Randall Division of Cell and Molecular Biophysics, King's College London, London SE1 1UL, United Kingdom; Medical Research Council and Asthma UK Centre in Allergic Mechanisms of Asthma, London SE1 9RT, United Kingdom. Electronic address: anna.davies@kcl.ac.uk.
Organizational Affiliation: