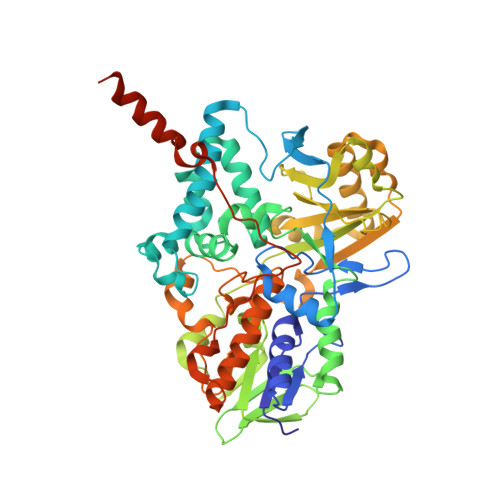
Kinetic and Structural Analysis of the Irreversible Inhibition of Human Monoamine Oxidases by Ass234, a Multi-Target Compound Designed for Use in Alzheimer'S Disease.
Esteban, G., Allan, J., Samadi, A., Mattevi, A., Unzeta, M., Marco-Contelles, J., Binda, C., Ramsay, R.R.(2014) Biochim Biophys Acta 1844: 1104
- PubMed: 24642166 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2014.03.006
- Primary Citation Related Structures:
4CRT - PubMed Abstract:
Monoamine oxidases (MAO) and cholinesterases are validated targets in the design of drugs for the treatment of Alzheimer's disease. The multi-target compound N-((5-(3-(1-benzylpiperidin-4-yl)propoxy)-1-methyl-1H-indol-2-yl)methyl)-N-methylprop-2-yn-1-amine (ASS234), bearing the MAO-inhibiting propargyl group attached to a donepezil moiety that inhibits cholinesterases, retained activity against human acetyl- and butyryl-cholinesterases. The inhibition of MAO A and MAO B by ASS234 was characterized and compared to other known MAO inhibitors. ASS234 was almost as effective as clorgyline (kinact/KI=3×10(6) min(-1)M(-1)) and was shown by structural studies to form the same N5 covalent adduct with the FAD cofactor.
- Departamento de Bioquímica i Biología Molecular, Institute of Neuroscience, Facultat de Medicina, Universitat Autònoma de Barcelona, 08193 Bellaterra, Barcelona, Spain.
Organizational Affiliation: