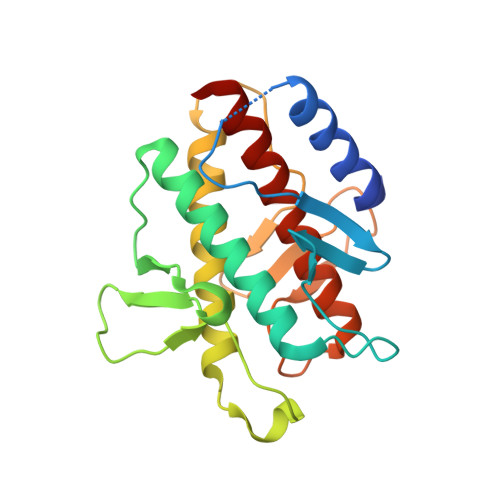
Crystal structure and biochemical analyses reveal Beclin 1 as a novel membrane binding protein
Huang, W., Choi, W., Hu, W., Mi, N., Guo, Q., Ma, M., Liu, M., Tian, Y., Lu, P., Wang, F.L., Deng, H., Liu, L., Gao, N., Yu, L., Shi, Y.(2012) Cell Res
- PubMed: 22310240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2012.24
- Primary Citation Related Structures:
4DDP - PubMed Abstract:
The Beclin 1 gene is a haplo-insufficient tumor suppressor and plays an essential role in autophagy. However, the molecular mechanism by which Beclin 1 functions remains largely unknown. Here we report the crystal structure of the evolutionarily conserved domain (ECD) of Beclin 1 at 1.6 Å resolution. Beclin 1 ECD exhibits a previously unreported fold, with three structural repeats arranged symmetrically around a central axis. Beclin 1 ECD defines a novel class of membrane-binding domain, with a strong preference for lipid membrane enriched with cardiolipin. The tip of a surface loop in Beclin 1 ECD, comprising three aromatic amino acids, acts as a hydrophobic finger to associate with lipid membrane, consequently resulting in the deformation of membrane and liposomes. Mutation of these aromatic residues rendered Beclin 1 unable to stably associate with lipid membrane in vitro and unable to fully rescue autophagy in Beclin 1-knockdown cells in vivo. These observations form an important framework for deciphering the biological functions of Beclin 1.
- Tsinghua-Peking Joint Center for Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: