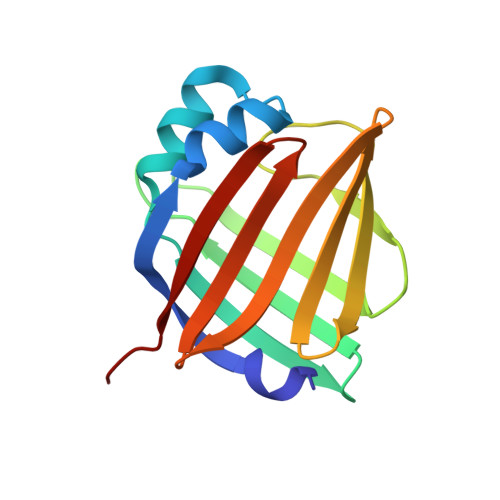
A bilirubin-inducible fluorescent protein from eel muscle
Kumagai, A., Ando, R., Miyatake, H., Greimel, P., Kobayashi, T., Hirabayashi, Y., Shimogori, T., Miyawaki, A.(2013) Cell 153: 1602-1611
- PubMed: 23768684 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2013.05.038
- Primary Citation Related Structures:
4I3B, 4I3C, 4I3D - PubMed Abstract:
The fluorescent protein toolbox has revolutionized experimental biology. Despite this advance, no fluorescent proteins have been identified from vertebrates, nor has chromogenic ligand-inducible activation or clinical utility been demonstrated. Here, we report the cloning and characterization of UnaG, a fluorescent protein from Japanese eel. UnaG belongs to the fatty-acid-binding protein (FABP) family, and expression in eel is restricted to small-diameter muscle fibers. On heterologous expression in cell lines or mouse brain, UnaG produces oxygen-independent green fluorescence. Remarkably, UnaG fluorescence is triggered by an endogenous ligand, bilirubin, a membrane-permeable heme metabolite and clinical health biomarker. The holoUnaG structure at 1.2 Å revealed a biplanar coordination of bilirubin by reversible π-conjugation, and we used this high-affinity and high-specificity interaction to establish a fluorescence-based human bilirubin assay with promising clinical utility. UnaG will be the prototype for a versatile class of ligand-activated fluorescent proteins, with applications in research, medicine, and bioengineering.
- Cell Function Dynamics, Brain Science Institute, RIKEN, 2-1 Hirosawa, Wako-city, Saitama 351-0198, Japan.
Organizational Affiliation: