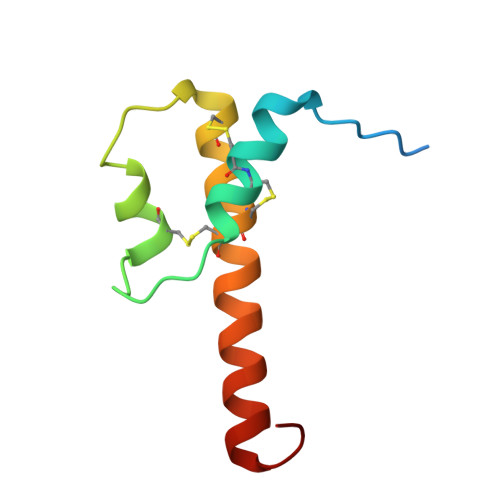
Efficient chemical synthesis of human complement protein C3a.
Ghassemian, A., Wang, C.I., Yau, M.K., Reid, R.C., Lewis, R.J., Fairlie, D.P., Alewood, P.F., Durek, T.(2013) Chem Commun (Camb) 49: 2356-2358
- PubMed: 23407800 Search on PubMed
- DOI: https://doi.org/10.1039/c3cc40537k
- Primary Citation Related Structures:
4I6O - PubMed Abstract:
We report the total chemical synthesis of human C3a by one-pot native chemical ligation of three unprotected peptide segments, followed by efficient in vitro folding that yielded the anaphylatoxin C3a in high yield and excellent purity. Synthetic C3a was fully active and its crystal structure at 2.1 Å resolution showed 3 helices and a C-terminal turn motif.
- Division of Chemistry and Structural Biology, Institute for Molecular Bioscience, The University of Queensland, St Lucia, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: