Design, synthesis and SAR of novel glucokinase activators.
Cheruvallath, Z.S., Gwaltney, S.L., Sabat, M., Tang, M., Feng, J., Wang, H., Miura, J., Guntupalli, P., Jennings, A., Hosfield, D., Lee, B., Wu, Y.(2013) Bioorg Med Chem Lett 23: 2166-2171
- PubMed: 23434031 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.01.093
- Primary Citation Related Structures:
4ISE, 4ISF, 4ISG - PubMed Abstract:
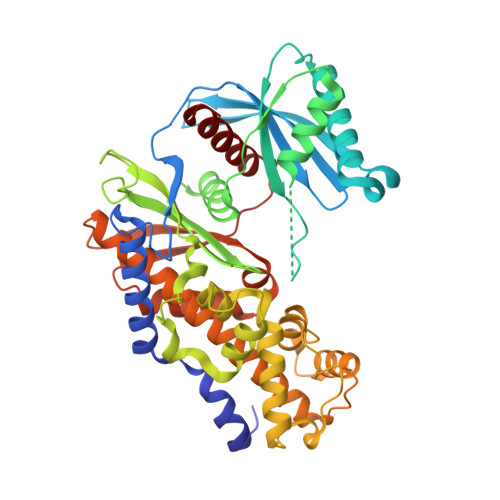
Guided by co-crystal structures of compounds 15, 22 and 30, an SBDD approach led to the discovery of the 6-methyl pyridone series as a novel class of GKAs that potently activate GK in enzyme and cell assays. Anti-diabetic OGTT efficacy was demonstrated with 54 in a mouse model of type 2 diabetes.
- Takeda California, 10410 Science Center Drive, San Diego, CA 92121, USA.
Organizational Affiliation: