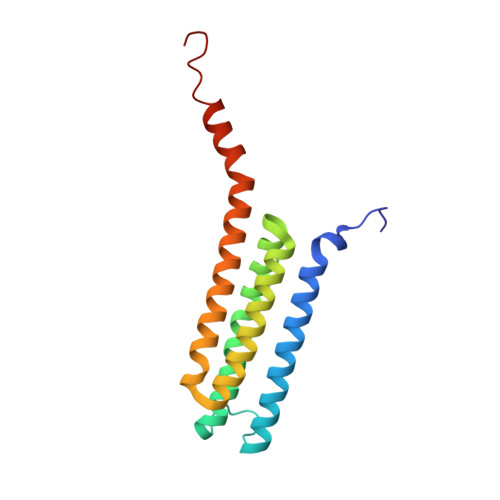
Crystal Structures of Leukotriene C4 Synthase in Complex with Product Analogs: IMPLICATIONS FOR THE ENZYME MECHANISM.
Niegowski, D., Kleinschmidt, T., Olsson, U., Ahmad, S., Rinaldo-Matthis, A., Haeggstrom, J.Z.(2014) J Biological Chem 289: 5199-5207
- PubMed: 24366866 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.534628
- Primary Citation Related Structures:
4J7T, 4J7Y, 4JC7, 4JCZ, 4JRZ - PubMed Abstract:
Leukotriene (LT) C4 synthase (LTC4S) catalyzes the conjugation of the fatty acid LTA4 with the tripeptide GSH to produce LTC4, the parent compound of the cysteinyl leukotrienes, important mediators of asthma. Here we mutated Trp-116 in human LTC4S, a residue proposed to play a key role in substrate binding, into an Ala or Phe. Biochemical and structural characterization of these mutants along with crystal structures of the wild type and mutated enzymes in complex with three product analogs, viz. S-hexyl-, 4-phenyl-butyl-, and 2-hydroxy-4-phenyl-butyl-glutathione, provide new insights to binding of substrates and product, identify a new conformation of the GSH moiety at the active site, and suggest a route for product release, aided by Trp-116.
- From the Division of Chemistry II, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-107 77 Stockholm, Sweden.
Organizational Affiliation: