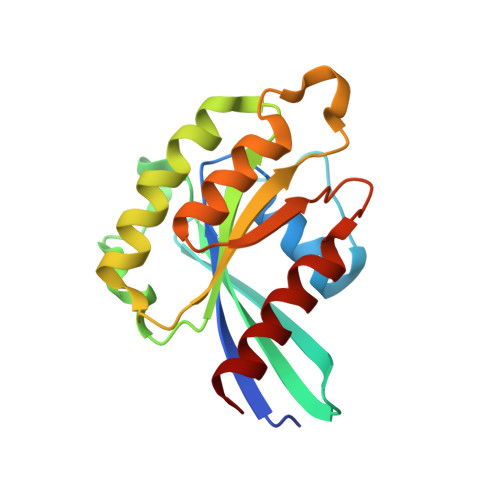
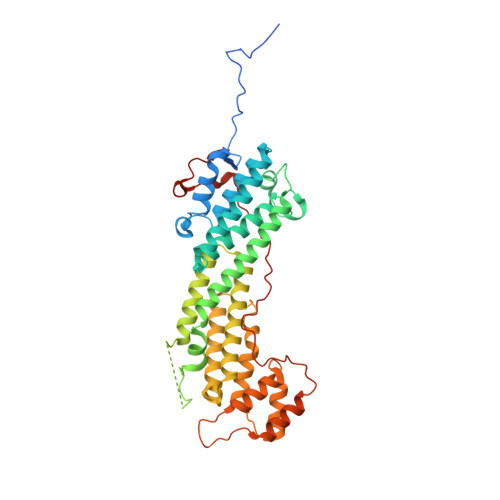
Structural basis of myosin V Rab GTPase-dependent cargo recognition.
Pylypenko, O., Attanda, W., Gauquelin, C., Lahmani, M., Coulibaly, D., Baron, B., Hoos, S., Titus, M.A., England, P., Houdusse, A.M.(2013) Proc Natl Acad Sci U S A 110: 20443-20448
- PubMed: 24248336
- DOI: https://doi.org/10.1073/pnas.1314329110
- Primary Citation Related Structures:
4LWZ, 4LX0, 4LX1, 4LX2 - PubMed Abstract:
Specific recognition of the cargo that molecular motors transport or tether to cytoskeleton tracks allows them to perform precise cellular functions at particular times and positions in cells. However, very little is known about how evolution has favored conservation of functions for some isoforms, while also allowing for the generation of new recognition sites and specialized cellular functions. Here we present several crystal structures of the myosin Va or the myosin Vb globular tail domain (GTD) that gives insights into how the motor is linked to the recycling membrane compartments via Rab11 or to the melanosome membrane via recognition of the melanophilin adaptor that binds to Rab27a. The structures illustrate how the Rab11-binding site has been conserved during evolution and how divergence at another site of the GTD allows more specific interactions such as the specific recognition of melanophilin by the myosin Va isoform. With atomic structural insights, these structures also show how either the partner or the GTD structural plasticity upon association is critical for selective recruitment of the motor.
- Structural Motility, Institut Curie, 75248 Paris Cedex 05, France.
Organizational Affiliation: