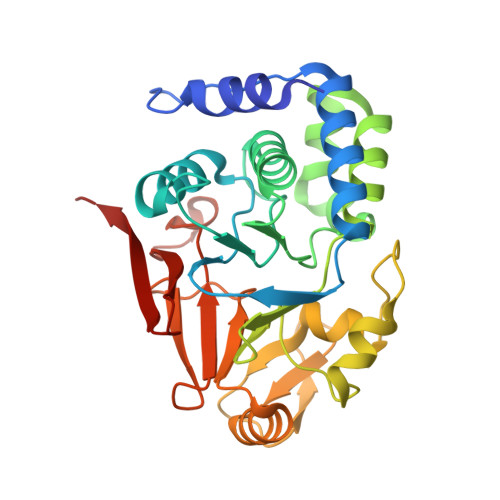
Understanding the antagonism of retinoblastoma protein dephosphorylation by PNUTS provides insights into the PP1 regulatory code.
Choy, M.S., Hieke, M., Kumar, G.S., Lewis, G.R., Gonzalez-Dewhitt, K.R., Kessler, R.P., Stein, B.J., Hessenberger, M., Nairn, A.C., Peti, W., Page, R.(2014) Proc Natl Acad Sci U S A 111: 4097-4102
- PubMed: 24591642
- DOI: https://doi.org/10.1073/pnas.1317395111
- Primary Citation of Related Structures:
4MOV, 4MOY, 4MP0 - PubMed Abstract:
The serine/threonine protein phosphatase 1 (PP1) dephosphorylates hundreds of key biological targets by associating with nearly 200 regulatory proteins to form highly specific holoenzymes. However, how these proteins direct PP1 specificity and the ability to predict how these PP1 interacting proteins bind PP1 from sequence alone is still missing. PP1 nuclear targeting subunit (PNUTS) is a PP1 targeting protein that, with PP1, plays a central role in the nucleus, where it regulates chromatin decondensation, RNA processing, and the phosphorylation state of fundamental cell cycle proteins, including the retinoblastoma protein (Rb), p53, and MDM2. The molecular function of PNUTS in these processes is completely unknown. Here, we show that PNUTS, which is intrinsically disordered in its free form, interacts strongly with PP1 in a highly extended manner. Unexpectedly, PNUTS blocks one of PP1's substrate binding grooves while leaving the active site accessible. This interaction site, which we have named the arginine site, allowed us to define unique PP1 binding motifs, which advances our ability to predict how more than a quarter of the known PP1 regulators bind PP1. Additionally, the structure shows how PNUTS inhibits the PP1-mediated dephosphorylation of critical substrates, especially Rb, by blocking their binding sites on PP1, insights that are providing strategies for selectively enhancing Rb activity.
- Departments of Molecular Pharmacology, Physiology, and Biotechnology, Molecular Biology, Cell Biology, and Biochemistry, and Chemistry, Brown University, Providence, RI 02912.
Organizational Affiliation: