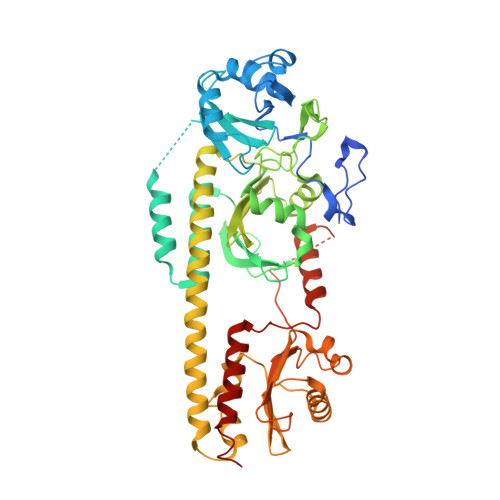
Signal amplification and transduction in phytochrome photosensors
Takala, H., Bjorling, A., Berntsson, O., Lehtivuori, H., Niebling, S., Hoernke, M., Kosheleva, I., Henning, R., Menzel, A., Ihalainen, J.A., Westenhoff, S.(2014) Nature 509: 245-248
- PubMed: 24776794
- DOI: https://doi.org/10.1038/nature13310
- Primary Citation of Related Structures:
4O01, 4O0P - PubMed Abstract:
Sensory proteins must relay structural signals from the sensory site over large distances to regulatory output domains. Phytochromes are a major family of red-light-sensing kinases that control diverse cellular functions in plants, bacteria and fungi. Bacterial phytochromes consist of a photosensory core and a carboxy-terminal regulatory domain. Structures of photosensory cores are reported in the resting state and conformational responses to light activation have been proposed in the vicinity of the chromophore. However, the structure of the signalling state and the mechanism of downstream signal relay through the photosensory core remain elusive. Here we report crystal and solution structures of the resting and activated states of the photosensory core of the bacteriophytochrome from Deinococcus radiodurans. The structures show an open and closed form of the dimeric protein for the activated and resting states, respectively. This nanometre-scale rearrangement is controlled by refolding of an evolutionarily conserved 'tongue', which is in contact with the chromophore. The findings reveal an unusual mechanism in which atomic-scale conformational changes around the chromophore are first amplified into an ångstrom-scale distance change in the tongue, and further grow into a nanometre-scale conformational signal. The structural mechanism is a blueprint for understanding how phytochromes connect to the cellular signalling network.
- Nanoscience Center, Department of Biological and Environmental Science, University of Jyväskylä, 40014 Jyväskylä, Finland.
Organizational Affiliation: