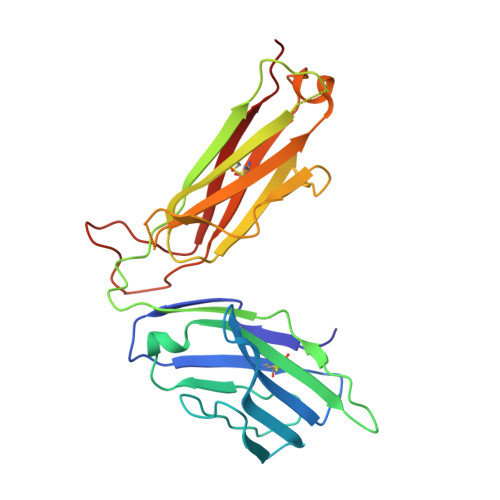
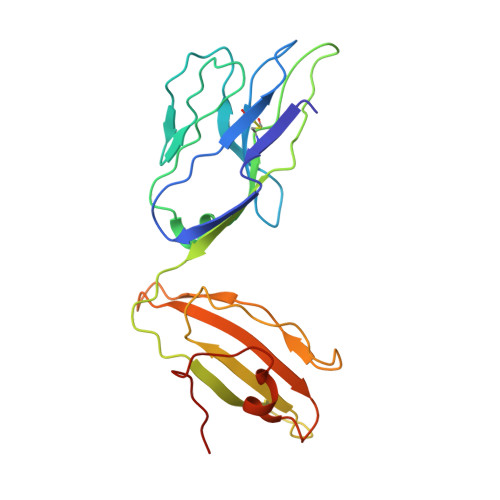
Molecular basis of mycobacterial lipid antigen presentation by CD1c and its recognition by alpha beta T cells.
Roy, S., Ly, D., Li, N.S., Altman, J.D., Piccirilli, J.A., Moody, D.B., Adams, E.J.(2014) Proc Natl Acad Sci U S A 111: E4648-E4657
- PubMed: 25298532 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1408549111
- Primary Citation Related Structures:
4ONH, 4ONO - PubMed Abstract:
CD1c is a member of the group 1 CD1 family of proteins that are specialized for lipid antigen presentation. Despite high cell surface expression of CD1c on key antigen-presenting cells and the discovery of its mycobacterial lipid antigen presentation capability, the molecular basis of CD1c recognition by T cells is unknown. Here we present a comprehensive functional and molecular analysis of αβ T-cell receptor (TCR) recognition of CD1c presenting mycobacterial phosphomycoketide antigens. Our structure of CD1c with the mycobacterial phosphomycoketide (PM) shows similarities to that of CD1c-mannosyl-β1-phosphomycoketide in that the A' pocket accommodates the mycoketide alkyl chain; however, the phosphate head-group of PM is shifted ∼6 Å in relation to that of mannosyl-β1-PM. We also demonstrate a bona fide interaction between six human TCRs and CD1c-mycoketide complexes, measuring high to moderate affinities. The crystal structure of the DN6 TCR and mutagenic studies reveal a requirement of five complementarity determining region (CDR) loops for CD1c recognition. Furthermore, mutagenesis of CD1c reveals residues in both the α1 and α2 helices involved in TCR recognition, yet not entirely overlapping among the examined TCRs. Unlike patterns for MHC I, no archetypical binding footprint is predicted to be shared by CD1c-reactive TCRs, even when recognizing the same or similar antigens.
- Departments of Biochemistry and Molecular Biology and.
Organizational Affiliation: