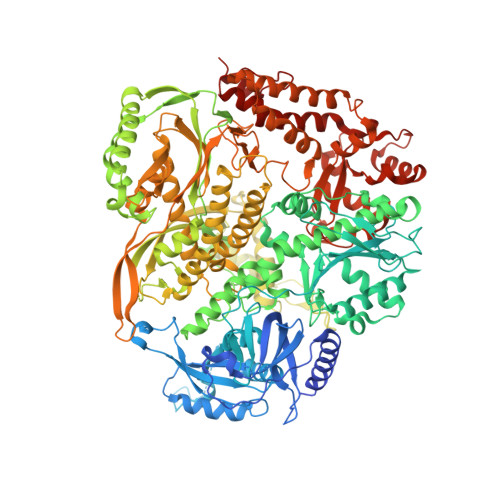
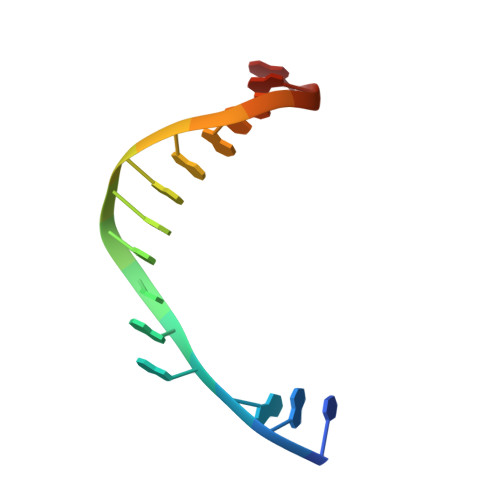
Crystal Structure of Yeast DNA Polymerase epsilon Catalytic Domain.
Jain, R., Rajashankar, K.R., Buku, A., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2014) PLoS One 9: e94835-e94835
- PubMed: 24733111 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0094835
- Primary Citation Related Structures:
4PTF - PubMed Abstract:
DNA polymerase ε (Polε) is a multi-subunit polymerase that contributes to genomic stability via its roles in leading strand replication and the repair of damaged DNA. Here we report the ternary structure of the Polε catalytic subunit (Pol2) bound to a nascent G:C base pair (Pol2G:C). Pol2G:C has a typical B-family polymerase fold and embraces the template-primer duplex with the palm, fingers, thumb and exonuclease domains. The overall arrangement of domains is similar to the structure of Pol2T:A reported recently, but there are notable differences in their polymerase and exonuclease active sites. In particular, we observe Ca2+ ions at both positions A and B in the polymerase active site and also observe a Ca2+ at position B of the exonuclease site. We find that the contacts to the nascent G:C base pair in the Pol2G:C structure are maintained in the Pol2T:A structure and reflect the comparable fidelity of Pol2 for nascent purine-pyrimidine and pyrimidine-purine base pairs. We note that unlike that of Pol3, the shape of the nascent base pair binding pocket in Pol2 is modulated from the major grove side by the presence of Tyr431. Together with Pol2T:A, our results provide a framework for understanding the structural basis of high fidelity DNA synthesis by Pol2.
- Department of Structural and Chemical Biology, Mount Sinai School of Medicine, New York, New York, United States of America.
Organizational Affiliation: