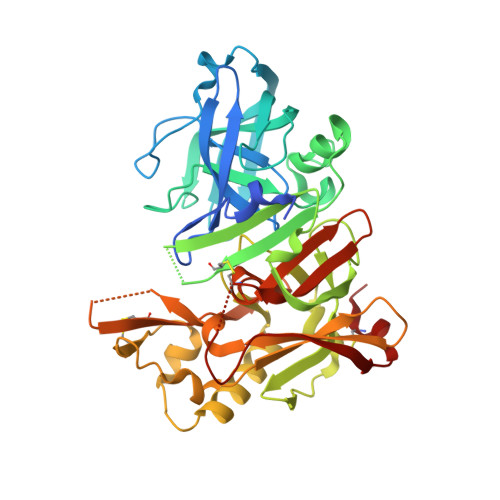
Methyl-substitution of an iminohydantoin spiropiperidine beta-secretase (BACE-1) inhibitor has a profound effect on its potency.
Egbertson, M., McGaughey, G.B., Pitzenberger, S.M., Stauffer, S.R., Coburn, C.A., Stachel, S.J., Yang, W., Barrow, J.C., Neilson, L.A., McWherter, M., Perlow, D., Fahr, B., Munshi, S., Allison, T.J., Holloway, K., Selnick, H.G., Yang, Z., Swestock, J., Simon, A.J., Sankaranarayanan, S., Colussi, D., Tugusheva, K., Lai, M.T., Pietrak, B., Haugabook, S., Jin, L., Chen, I.W., Holahan, M., Stranieri-Michener, M., Cook, J.J., Vacca, J., Graham, S.L.(2015) Bioorg Med Chem Lett 25: 4812-4819
- PubMed: 26195137
- DOI: https://doi.org/10.1016/j.bmcl.2015.06.082
- Primary Citation Related Structures:
4ZPE, 4ZPF, 4ZPG - PubMed Abstract:
The IC50 of a beta-secretase (BACE-1) lead compound was improved ∼200-fold from 11 μM to 55 nM through the addition of a single methyl group. Computational chemistry, small molecule NMR, and protein crystallography capabilities were used to compare the solution conformation of the ligand under varying pH conditions to its conformation when bound in the active site. Chemical modification then explored available binding pockets adjacent to the ligand. A strategically placed methyl group not only maintained the required pKa of the piperidine nitrogen and filled a small hydrophobic pocket, but more importantly, stabilized the conformation best suited for optimized binding to the receptor.
- Medicinal Chemistry Department, WP14-2 Merck and Co., West Point, PA 19486, USA. Electronic address: melissaegbertson@gmail.com.
Organizational Affiliation: