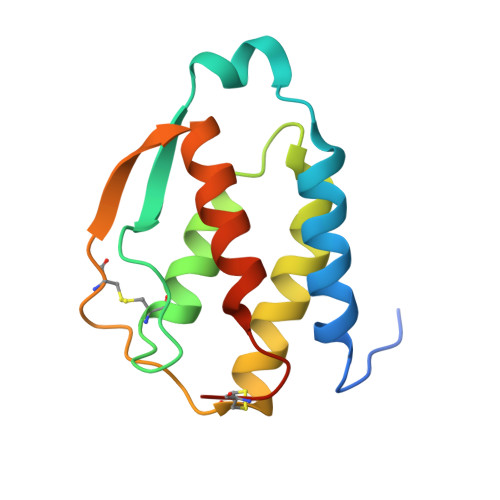
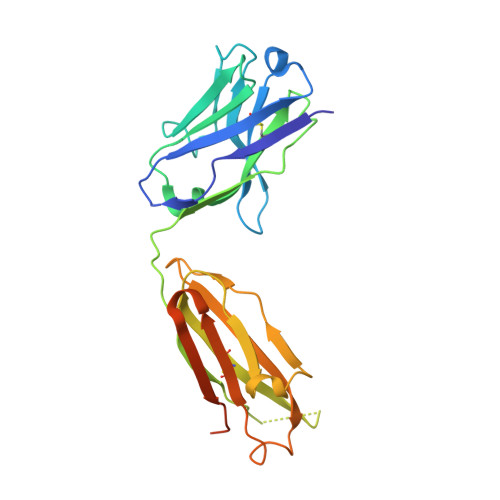
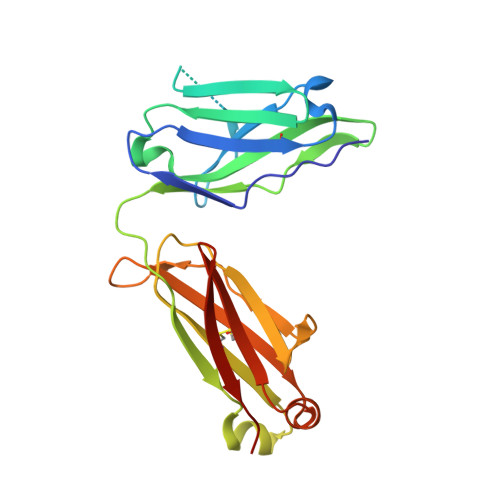
Molecular basis of in vitro affinity maturation and functional evolution of a neutralizing anti-human GM-CSF antibody.
Eylenstein, R., Weinfurtner, D., Hartle, S., Strohner, R., Bottcher, J., Augustin, M., Ostendorp, R., Steidl, S.(2016) MAbs 8: 176-186
- PubMed: 26406987
- DOI: https://doi.org/10.1080/19420862.2015.1099774
- Primary Citation Related Structures:
5C7X, 5D70, 5D71, 5D72, 5D7S - PubMed Abstract:
X-ray structure analysis of 4 antibody Fab fragments, each in complex with human granulocyte macrophage colony stimulating factor (GM-CSF), was performed to investigate the changes at the protein-protein binding interface during the course of in vitro affinity maturation by phage display selection. The parental antibody MOR03929 was compared to its derivatives MOR04252 (CDR-H2 optimized), MOR04302 (CDR-L3 optimized) and MOR04357 (CDR-H2 and CDR-L3 optimized). All antibodies bind to a conformational epitope that can be divided into 3 sub-epitopes. Specifically, MOR04357 binds to a region close to the GM-CSF N-terminus (residues 11-24), a short second sub-epitope (residues 83-89) and a third at the C-terminus (residues 112-123). Modifications introduced during affinity maturation in CDR-H2 and CDR-L3 led to the establishment of additional hydrogen bonds and van der Waals contacts, respectively, providing a rationale for the observed improvement in binding affinity and neutralization potency. Once GM-CSF is complexed to the antibodies, modeling predicts a sterical clash with GM-CSF binding to GM-CSF receptor α and β chain. This predicted mutually exclusive binding was confirmed by a GM-CSF receptor α chain ligand binding inhibition assay. Finally, high throughput sequencing of clones obtained after affinity maturation phage display pannings revealed highly selected consensus sequences for CDR-H2 as well for CDR-L3, which are in accordance with the sequence of the highest affinity antibody MOR04357. The resolved crystal structures highlight the criticality of these strongly selected residues for high affinity interaction with GM-CSF.
- a MorphoSys AG ; Lena-Christ-Str. 48; 82152 Martinsried ; Germany.
Organizational Affiliation: