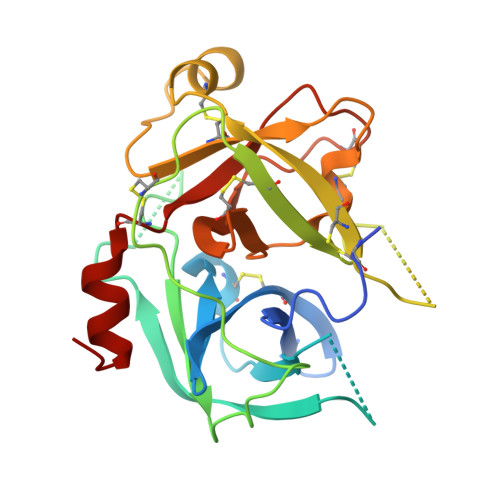
Structural basis for the Zn2+ inhibition of the zymogen-like kallikrein-related peptidase 10.
Debela, M., Magdolen, V., Bode, W., Brandstetter, H., Goettig, P.(2016) Biol Chem 397: 1251-1264
- PubMed: 27611765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1515/hsz-2016-0205
- Primary Citation Related Structures:
5LPE, 5LPF - PubMed Abstract:
Although kallikrein-related peptidase 10 (KLK10) is expressed in a variety of human tissues and body fluids, knowledge of its physiological functions is fragmentary. Similarly, the pathophysiology of KLK10 in cancer is not well understood. In some cancer types, a role as tumor suppressor has been suggested, while in others elevated expression is associated with poor patient prognosis. Active human KLK10 exhibits a unique, three residue longer N-terminus with respect to other serine proteases and an extended 99-loop nearly as long as in tissue kallikrein KLK1. Crystal structures of recombinant ligand-free KLK10 and a Zn2+ bound form explain to some extent the mixed trypsin- and chymotrypsin-like substrate specificity. Zn2+-inhibition of KLK10 appears to be based on a unique mechanism, which involves direct binding and blocking of the catalytic triad. Since the disordered N-terminus and several loops adopt a zymogen-like conformation, the active protease conformation is very likely induced by interaction with the substrate, in particular at the S1 subsite and at the unusual Ser193 as part of the oxyanion hole. The KLK10 structures indicate that the N-terminus, the nearby 75-, 148-, and the 99-loops are connected in an allosteric network, which is present in other trypsin-like serine proteases with several variations.