Inhibition of delta-secretase improves cognitive functions in mouse models of Alzheimer's disease.
Zhang, Z., Obianyo, O., Dall, E., Du, Y., Fu, H., Liu, X., Kang, S.S., Song, M., Yu, S.P., Cabrele, C., Schubert, M., Li, X., Wang, J.Z., Brandstetter, H., Ye, K.(2017) Nat Commun 8: 14740-14740
- PubMed: 28345579
- DOI: https://doi.org/10.1038/ncomms14740
- Primary Citation Related Structures:
5LU8, 5LU9, 5LUA, 5LUB - PubMed Abstract:
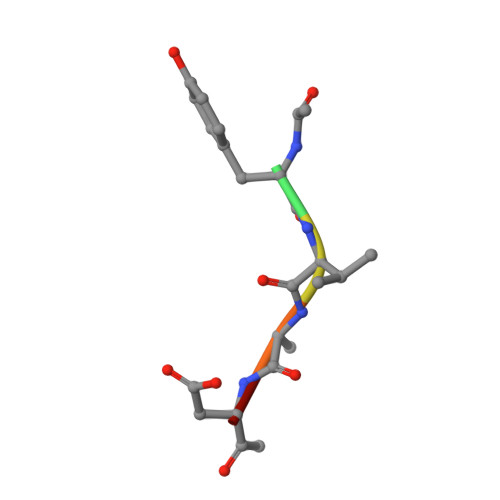
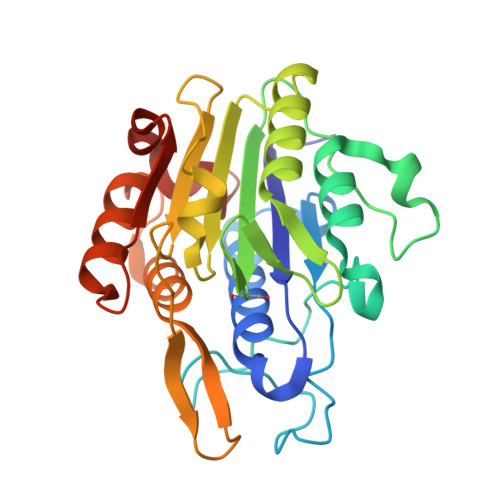
δ-secretase, also known as asparagine endopeptidase (AEP) or legumain, is a lysosomal cysteine protease that cleaves both amyloid precursor protein (APP) and tau, mediating the amyloid-β and tau pathology in Alzheimer's disease (AD). Here we report the therapeutic effect of an orally bioactive and brain permeable δ-secretase inhibitor in mouse models of AD. We performed a high-throughput screen and identified a non-toxic and selective δ-secretase inhibitor, termed compound 11, that specifically blocks δ-secretase but not other related cysteine proteases. Co-crystal structure analysis revealed a dual active site-directed and allosteric inhibition mode of this compound class. Chronic treatment of tau P301S and 5XFAD transgenic mice with this inhibitor reduces tau and APP cleavage, ameliorates synapse loss and augments long-term potentiation, resulting in protection of memory. Therefore, these findings demonstrate that this δ-secretase inhibitor may be an effective clinical therapeutic agent towards AD.
- Department of Pathology and Laboratory Medicine, Emory University School of Medicine, Atlanta, Georgia 30322, USA.
Organizational Affiliation: