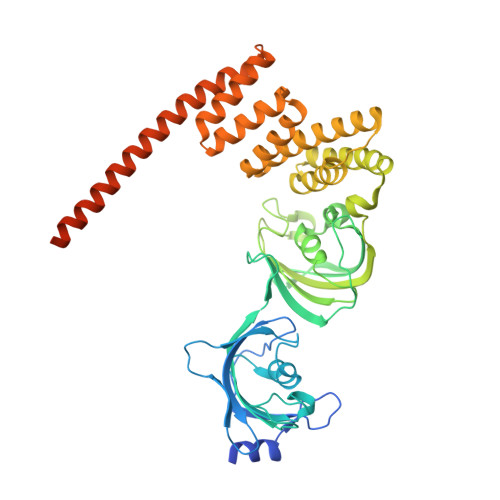
Combined x-ray crystallography and computational modeling approach to investigate the Hsp90 C-terminal peptide binding to FKBP51.
Kumar, R., Moche, M., Winblad, B., Pavlov, P.F.(2017) Sci Rep 7: 14288-14288
- PubMed: 29079741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-14731-z
- Primary Citation Related Structures:
5NJX, 5OMP - PubMed Abstract:
FK506 binding protein of 51 kDa (FKBP51) is a heat shock protein 90 (Hsp90) co-chaperone involved in the regulation of steroid hormone receptors activity. It is known for its role in various regulatory pathways implicated in mood and stress-related disorders, cancer, obesity, Alzheimer's disease and corticosteroid resistant asthma. It consists of two FKBP12 like active peptidyl prolyl isomerase (PPIase) domains (an active FK1 and inactive FK2 domain) and one tetratricopeptide repeat (TPR) domain that mediates interaction with Hsp90 via its C-terminal MEEVD peptide. Here, we report a combined x-ray crystallography and molecular dynamics study to reveal the binding mechanism of Hsp90 MEEVD peptide to the TPR domain of FKBP51. The results demonstrated that the Hsp90 C-terminal peptide binds to the TPR domain of FKBP51 with the help of di-carboxylate clamp involving Lys272, Glu273, Lys352, Asn322, and Lys329 which are conserved throughout several di-carboxylate clamp TPR proteins. Interestingly, the results from molecular dynamics study are also in agreement to the complex structure where all the contacts between these two partners were consistent throughout the simulation period. In a nutshell, our findings provide new opportunity to engage this important protein-protein interaction target by small molecules designed by structure based drug design strategy.
- Department of Neurobiology, Care Sciences and Society, Center for Alzheimer Research, Division of Neurogeriatrics, Karolinska Institutet, Huddinge, Sweden.
Organizational Affiliation: