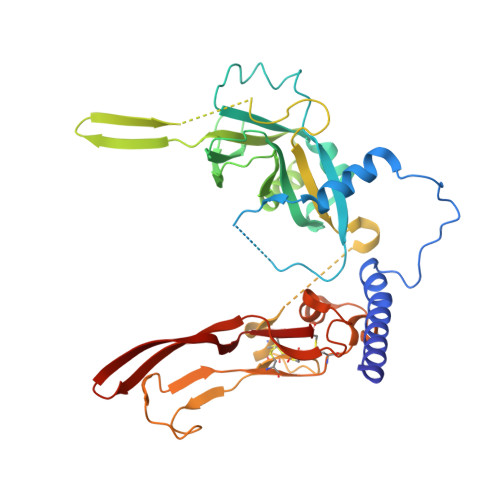
Prodomain-growth factor swapping in the structure of pro-TGF-beta 1.
Zhao, B., Xu, S., Dong, X., Lu, C., Springer, T.A.(2018) J Biological Chem 293: 1579-1589
- PubMed: 29109152
- DOI: https://doi.org/10.1074/jbc.M117.809657
- Primary Citation Related Structures:
5VQF, 5VQP - PubMed Abstract:
TGF-β is synthesized as a proprotein that dimerizes in the endoplasmic reticulum. After processing in the Golgi to cleave the N-terminal prodomain from the C-terminal growth factor (GF) domain in each monomer, pro-TGF-β is secreted and stored in latent complexes. It is unclear which prodomain and GF monomer are linked before proprotein convertase cleavage and how much conformational change occurs following cleavage. We have determined a structure of pro-TGF-β1 with the proprotein convertase cleavage site mutated to mimic the structure of the TGF-β1 proprotein. Structure, mutation, and model building demonstrate that the prodomain arm domain in one monomer is linked to the GF that interacts with the arm domain in the other monomer in the dimeric structure ( i.e. the prodomain arm domain and GF domain in each monomer are swapped). Swapping has important implications for the mechanism of biosynthesis in the TGF-β family and is relevant to the mechanism for preferential formation of heterodimers over homodimers for some members of the TGF-β family. Our structure, together with two previous ones, also provides insights into which regions of the prodomain-GF complex are highly structurally conserved and which are perturbed by crystal lattice contacts.
- From the Children's Hospital Boston and Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115 and.
Organizational Affiliation: