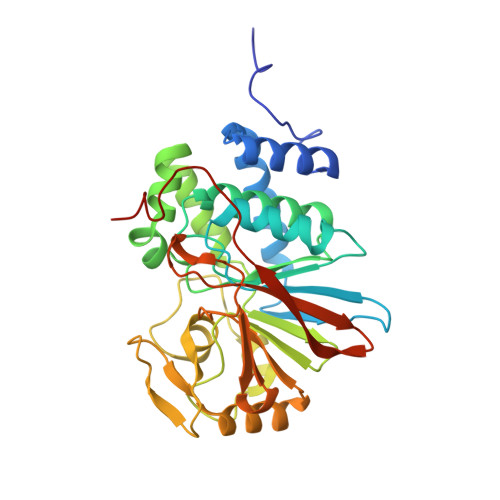
The Antitumor Drug LB-100 Is a Catalytic Inhibitor of Protein Phosphatase 2A (PPP2CA) and 5 (PPP5C) Coordinating with the Active-Site Catalytic Metals in PPP5C.
D'Arcy, B.M., Swingle, M.R., Papke, C.M., Abney, K.A., Bouska, E.S., Prakash, A., Honkanen, R.E.(2019) Mol Cancer Ther 18: 556-566
- PubMed: 30679389
- DOI: https://doi.org/10.1158/1535-7163.MCT-17-1143
- Primary Citation Related Structures:
5WG8 - PubMed Abstract:
LB-100 is an experimental cancer therapeutic with cytotoxic activity against cancer cells in culture and antitumor activity in animals. The first phase I trial (NCT01837667) evaluating LB-100 recently concluded that safety and efficacy parameters are favorable for further clinical testing. Although LB-100 is widely reported as a specific inhibitor of serine/threonine phosphatase 2A (PP2AC/ PPP2CA:PPP2CB ), we could find no experimental evidence in the published literature demonstrating the specific engagement of LB-100 with PP2A in vitro , in cultured cells, or in animals. Rather, the premise for LB-100 targeting PP2AC is derived from studies that measure phosphate released from a phosphopeptide (K-R-pT-I-R-R) or inferred from the ability of LB-100 to mimic activity previously reported to result from the inhibition of PP2AC by other means. PP2AC and PPP5C share a common catalytic mechanism. Here, we demonstrate that the phosphopeptide used to ascribe LB-100 specificity for PP2A is also a substrate for PPP5C. Inhibition assays using purified enzymes demonstrate that LB-100 is a catalytic inhibitor of both PP2AC and PPP5C. The structure of PPP5C cocrystallized with LB-100 was solved to a resolution of 1.65Å, revealing that the 7-oxabicyclo[2.2.1]heptane-2,3-dicarbonyl moiety coordinates with the metal ions and key residues that are conserved in both PP2AC and PPP5C. Cell-based studies revealed some known actions of LB-100 are mimicked by the genetic disruption of PPP5C These data demonstrate that LB-100 is a catalytic inhibitor of both PP2AC and PPP5C and suggest that the observed antitumor activity might be due to an additive effect achieved by suppressing both PP2A and PPP5C.
- USA Mitchell Cancer Institute, Mobile, Alabama.
Organizational Affiliation: