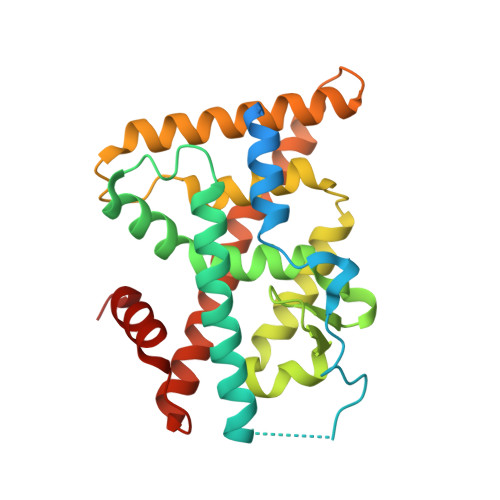
PGE1 and PGA1 bind to Nurr1 and activate its transcriptional function.
Rajan, S., Jang, Y., Kim, C.H., Kim, W., Toh, H.T., Jeon, J., Song, B., Serra, A., Lescar, J., Yoo, J.Y., Beldar, S., Ye, H., Kang, C., Liu, X.W., Feitosa, M., Kim, Y., Hwang, D., Goh, G., Lim, K.L., Park, H.M., Lee, C.H., Oh, S.F., Petsko, G.A., Yoon, H.S., Kim, K.S.(2020) Nat Chem Biol
- PubMed: 32451509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-020-0553-6
- Primary Citation Related Structures:
5Y41 - PubMed Abstract:
The orphan nuclear receptor Nurr1 is critical for the development, maintenance and protection of midbrain dopaminergic (mDA) neurons. Here we show that prostaglandin E1 (PGE1) and its dehydrated metabolite, PGA1, directly interact with the ligand-binding domain (LBD) of Nurr1 and stimulate its transcriptional function. We also report the crystallographic structure of Nurr1-LBD bound to PGA1 at 2.05 Å resolution. PGA1 couples covalently to Nurr1-LBD by forming a Michael adduct with Cys566, and induces notable conformational changes, including a 21° shift of the activation function-2 helix (H12) away from the protein core. Furthermore, PGE1/PGA1 exhibit neuroprotective effects in a Nurr1-dependent manner, prominently enhance expression of Nurr1 target genes in mDA neurons and improve motor deficits in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine-lesioned mouse models of Parkinson's disease. Based on these results, we propose that PGE1/PGA1 represent native ligands of Nurr1 and can exert neuroprotective effects on mDA neurons, via activation of Nurr1's transcriptional function.
- School of Biological Sciences, Nanyang Technological University, Singapore, Singapore.
Organizational Affiliation: