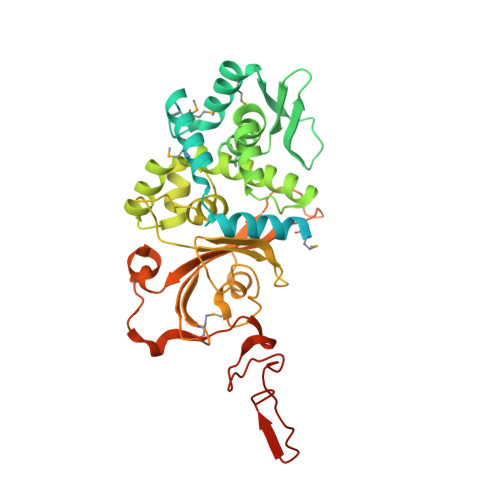
Structural and functional insight into the Mycobacterium tuberculosis protein PrpR reveals a novel type of transcription factor.
Tang, S., Hicks, N.D., Cheng, Y.S., Silva, A., Fortune, S.M., Sacchettini, J.C.(2019) Nucleic Acids Res 47: 9934-9949
- PubMed: 31504787
- DOI: https://doi.org/10.1093/nar/gkz724
- Primary Citation of Related Structures:
6CYJ, 6CYY, 6CZ6, 6D2S - PubMed Abstract:
The pathogenicity of Mycobacterium tuberculosis depends upon its ability to catabolize host cholesterol. Upregulation of the methylcitrate cycle (MCC) is required to assimilate and detoxify propionyl-CoA, a cholesterol degradation product. The transcription of key genes prpC and prpD in MCC is activated by MtPrpR, a member of a family of prokaryotic transcription factors whose structures and modes of action have not been clearly defined. We show that MtPrpR has a novel overall structure and directly binds to CoA or short-chain acyl-CoA derivatives to form a homotetramer that covers the binding cavity and locks CoA tightly inside the protein. The regulation of this process involves a [4Fe4S] cluster located close to the CoA-binding cavity on a neighboring chain. Mutations in the [4Fe4S] cluster binding residues rendered MtPrpR incapable of regulating MCC gene transcription. The structure of MtPrpR without the [4Fe4S] cluster-binding region shows a conformational change that prohibits CoA binding. The stability of this cluster means it is unlikely a redox sensor but may function by sensing ambient iron levels. These results provide mechanistic insights into this family of critical transcription factors who share similar structures and regulate gene transcription using a combination of acyl-CoAs and [4Fe4S] cluster.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77840, USA.
Organizational Affiliation: