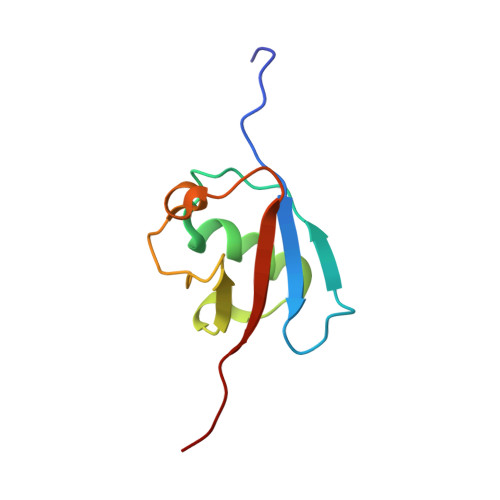
The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation.
Aichem, A., Anders, S., Catone, N., Stotz, S., Berg, A., Schwab, R., Scheuermann, S., Bialas, J., Schutz-Stoffregen, M.C., Schmidtke, G., Peter, C., Groettrup, M., Wiesner, S.(2018) Nat Commun 9: 3321-3321
- PubMed: 30127417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-05776-3
- Primary Citation Related Structures:
6GF1, 6GF2 - PubMed Abstract:
FAT10 is a ubiquitin-like modifier that directly targets proteins for proteasomal degradation. Here, we report the high-resolution structures of the two individual ubiquitin-like domains (UBD) of FAT10 that are joined by a flexible linker. While the UBDs of FAT10 show the typical ubiquitin-fold, their surfaces are entirely different from each other and from ubiquitin explaining their unique binding specificities. Deletion of the linker abrogates FAT10-conjugation while its mutation blocks auto-FAT10ylation of the FAT10-conjugating enzyme USE1 but not bulk conjugate formation. FAT10- but not ubiquitin-mediated degradation is independent of the segregase VCP/p97 in the presence but not the absence of FAT10's unstructured N-terminal heptapeptide. Stabilization of the FAT10 UBDs strongly decelerates degradation suggesting that the intrinsic instability of FAT10 together with its disordered N-terminus enables the rapid, joint degradation of FAT10 and its substrates without the need for FAT10 de-conjugation and partial substrate unfolding.
- Division of Immunology, Department of Biology, University of Konstanz, Konstanz, D-78457, Germany.
Organizational Affiliation: