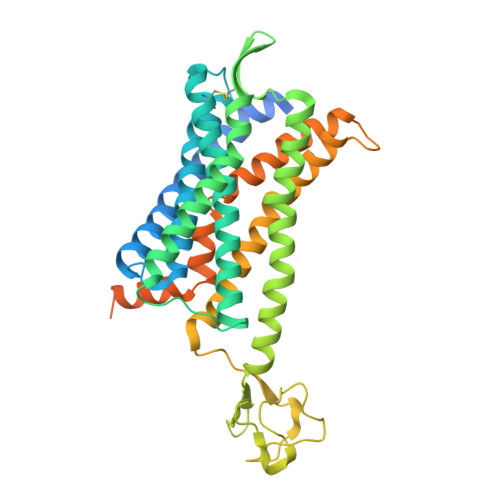
Crystal Structure of CC Chemokine Receptor 2A in Complex with an Orthosteric Antagonist Provides Insights for the Design of Selective Antagonists.
Apel, A.K., Cheng, R.K.Y., Tautermann, C.S., Brauchle, M., Huang, C.Y., Pautsch, A., Hennig, M., Nar, H., Schnapp, G.(2019) Structure 27: 427-438.e5
- PubMed: 30581043
- DOI: https://doi.org/10.1016/j.str.2018.10.027
- Primary Citation of Related Structures:
6GPS, 6GPX - PubMed Abstract:
We determined two crystal structures of the chemokine receptor CCR2A in complex with the orthosteric antagonist MK-0812. Full-length CCR2A, stabilized by rubredoxin and a series of five mutations were resolved at 3.3 Å. An N- and C-terminally truncated CCR2A construct was crystallized in an alternate crystal form, which yielded a 2.7 Å resolution structure using serial synchrotron crystallography. Our structures provide a clear structural explanation for the observed key role of residue E291 7.39 in high-affinity binding of several orthosteric CCR2 antagonists. By combining all the structural information collected, we generated models of co-structures for the structurally diverse pyrimidine amide class of CCR2 antagonists. Even though the representative Ex15 overlays well with MK-0812, it also interacts with the non-conserved H121 3.33 , resulting in a significant selectivity over CCR5. Insights derived from this work will facilitate drug discovery efforts directed toward highly selective CCR2 antagonists with potentially superior efficacy.
- Boehringer Ingelheim Pharma GmbH & Co. KG, Birkendorferstrasse 65, 88397 Biberach, Germany.
Organizational Affiliation: