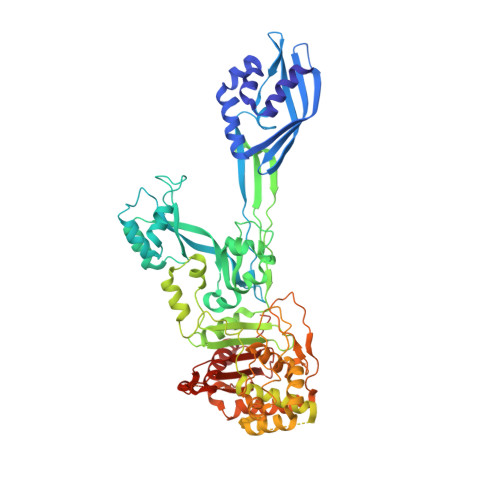
The Quinazolinone Allosteric Inhibitor of PBP 2a Synergizes with Piperacillin and Tazobactam against Methicillin-Resistant Staphylococcus aureus.
Janardhanan, J., Bouley, R., Martinez-Caballero, S., Peng, Z., Batuecas-Mordillo, M., Meisel, J.E., Ding, D., Schroeder, V.A., Wolter, W.R., Mahasenan, K.V., Hermoso, J.A., Mobashery, S., Chang, M.(2019) Antimicrob Agents Chemother 63
- PubMed: 30858202
- DOI: https://doi.org/10.1128/AAC.02637-18
- Primary Citation Related Structures:
6H5O, 6Q9N - PubMed Abstract:
The quinazolinones are a new class of antibacterials with in vivo efficacy against methicillin-resistant Staphylococcus aureus (MRSA). The quinazolinones target cell wall biosynthesis and have a unique mechanism of action by binding to the allosteric site of penicillin-binding protein 2a (PBP 2a). We investigated the potential for synergism of a lead quinazolinone with several antibiotics of different classes using checkerboard and time-kill assays. The quinazolinone synergized with β-lactam antibiotics. The combination of the quinazolinone with commercial piperacillin-tazobactam showed bactericidal synergy at sub-MICs of all three drugs. We demonstrated the efficacy of the triple-drug combination in a mouse MRSA neutropenic thigh infection model. The proposed mechanism for the synergistic activity in MRSA involves inhibition of the β-lactamase by tazobactam, which protects piperacillin from hydrolysis, which can then inhibit its target, PBP 2. Furthermore, the quinazolinone binds to the allosteric site of PBP 2a, triggering the allosteric response. This leads to the opening of the active site, which, in turn, binds another molecule of piperacillin. In other words, PBP 2a, which is not normally inhibited by piperacillin, becomes vulnerable to inhibition in the presence of the quinazolinone. The collective effect is the impairment of cell wall biosynthesis, with bactericidal consequence. Two crystal structures for complexes of the antibiotics with PBP 2a provide support for the proposed mechanism of action.
- Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, Indiana, USA.
Organizational Affiliation: