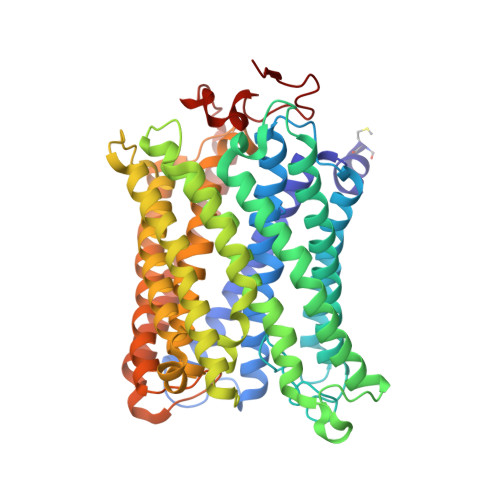
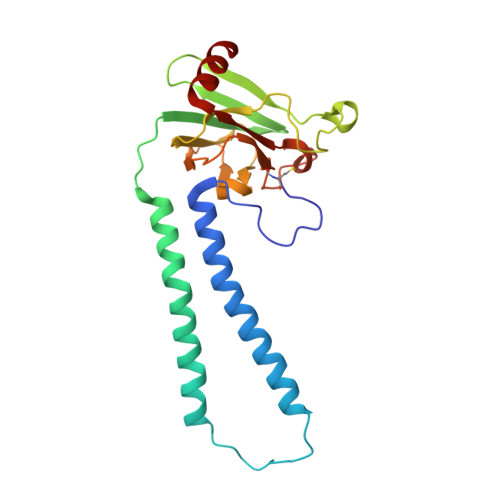
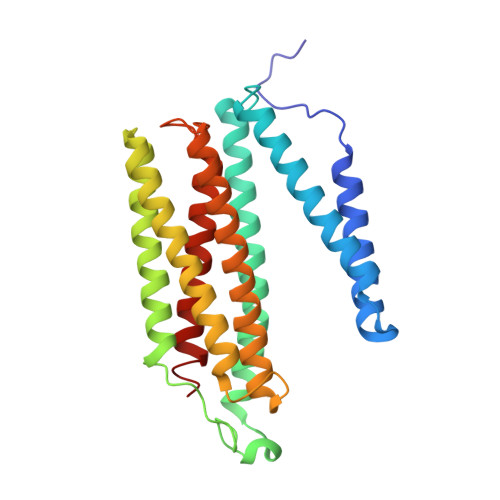
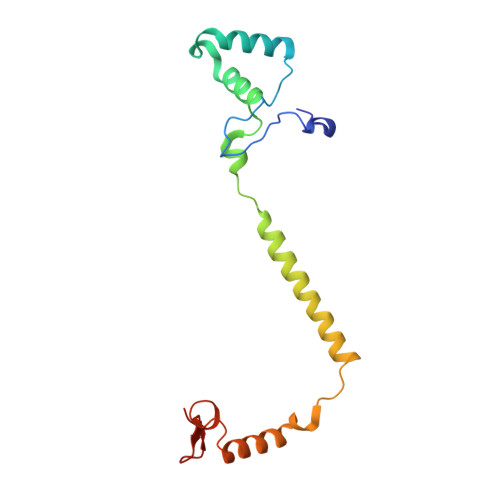
X-ray structures of catalytic intermediates of cytochromecoxidase provide insights into its O2activation and unidirectional proton-pump mechanisms.
Shimada, A., Etoh, Y., Kitoh-Fujisawa, R., Sasaki, A., Shinzawa-Itoh, K., Hiromoto, T., Yamashita, E., Muramoto, K., Tsukihara, T., Yoshikawa, S.(2020) J Biological Chem 295: 5818-5833
- PubMed: 32165497
- DOI: https://doi.org/10.1074/jbc.RA119.009596
- Primary Citation of Related Structures:
6JUW - PubMed Abstract:
Cytochrome c oxidase (CcO) reduces O 2 to water, coupled with a proton-pumping process. The structure of the O 2 -reduction site of CcO contains two reducing equivalents, Fe a 3 2+ and Cu B 1+ , and suggests that a peroxide-bound state (Fe a 3 3+ -O - -O - -Cu B 2+ ) rather than an O 2 -bound state (Fe a 3 2+ -O 2 ) is the initial catalytic intermediate. Unexpectedly, however, resonance Raman spectroscopy results have shown that the initial intermediate is Fe a 3 2+ -O 2 , whereas Fe a 3 3+ -O - -O - -Cu B 2+ is undetectable. Based on X-ray structures of static noncatalytic CcO forms and mutation analyses for bovine CcO, a proton-pumping mechanism has been proposed. It involves a proton-conducting pathway (the H-pathway) comprising a tandem hydrogen-bond network and a water channel located between the N- and P-side surfaces. However, a system for unidirectional proton-transport has not been experimentally identified. Here, an essentially identical X-ray structure for the two catalytic intermediates (P and F) of bovine CcO was determined at 1.8 Å resolution. A 1.70 Å Fe-O distance of the ferryl center could best be described as Fe a 3 4+ = O 2- , not as Fe a 3 4+ -OH - The distance suggests an ∼800-cm -1 Raman stretching band. We found an interstitial water molecule that could trigger a rapid proton-coupled electron transfer from tyrosine-OH to the slowly forming Fe a 3 3+ -O - -O - -Cu B 2+ state, preventing its detection, consistent with the unexpected Raman results. The H-pathway structures of both intermediates indicated that during proton-pumping from the hydrogen-bond network to the P-side, a transmembrane helix closes the water channel connecting the N-side with the hydrogen-bond network, facilitating unidirectional proton-pumping during the P-to-F transition.
- Picobiology Institute, Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori, Akoh, Hyogo 678-1297, Japan.
Organizational Affiliation: