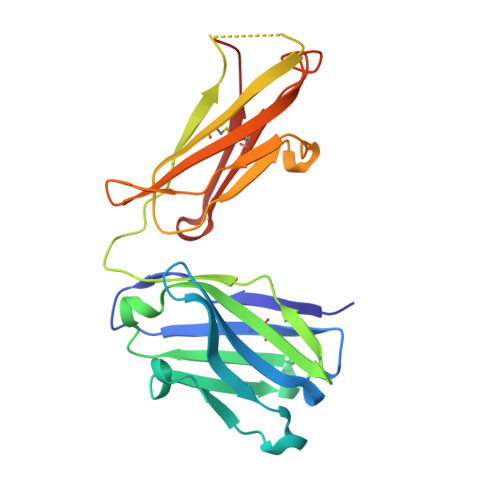
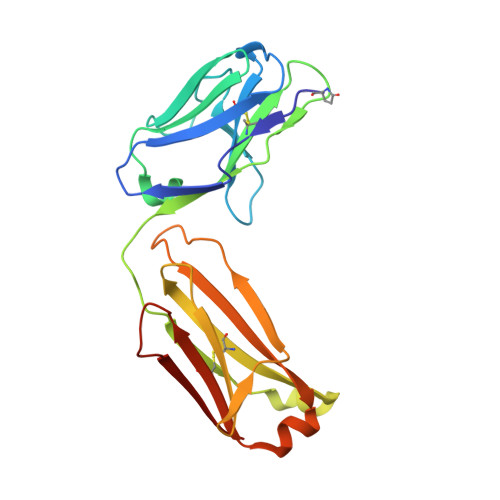
Structural basis of polyethylene glycol recognition by antibody.
Lee, C.C., Su, Y.C., Ko, T.P., Lin, L.L., Yang, C.Y., Chang, S.S., Roffler, S.R., Wang, A.H.(2020) J Biomed Sci 27: 12-12
- PubMed: 31907057 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12929-019-0589-7
- Primary Citation Related Structures:
6JP7, 6JU0, 6JWC - PubMed Abstract:
Polyethylene glycol (PEG) is widely used in industry and medicine. Anti-PEG antibodies have been developed for characterizing PEGylated drugs and other applications. However, the underlying mechanism for specific PEG binding has not been elucidated. The Fab of two cognate anti-PEG antibodies 3.3 and 2B5 were each crystallized in complex with PEG, and their structures were determined by X-ray diffraction. The PEG-Fab interactions in these two crystals were analyzed and compared with those in a PEG-containing crystal of an unrelated anti-hemagglutinin 32D6-Fab. The PEG-binding stoichiometry was examined by using analytical ultracentrifuge (AUC). A common PEG-binding mode to 3.3 and 2B5 is seen with an S-shaped core PEG fragment bound to two dyad-related Fab molecules. A nearby satellite binding site may accommodate parts of a longer PEG molecule. The core PEG fragment mainly interacts with the heavy-chain residues D31, W33, L102, Y103 and Y104, making extensive contacts with the aromatic side chains. At the center of each half-circle of the S-shaped PEG, a water molecule makes alternating hydrogen bonds to the ether oxygen atoms, in a similar configuration to that of a crown ether-bound lysine. Each satellite fragment is clamped between two arginine residues, R52 from the heavy chain and R29 from the light chain, and also interacts with several aromatic side chains. In contrast, the non-specifically bound PEG fragments in the 32D6-Fab crystal are located in the elbow region or at lattice contacts. The AUC data suggest that 3.3-Fab exists as a monomer in PEG-free solution but forms a dimer in the presence of PEG-550-MME, which is about the size of the S-shaped core PEG fragment. The differing amino acids in 3.3 and 2B5 are not involved in PEG binding but engaged in dimer formation. In particular, the light-chain residue K53 of 2B5-Fab makes significant contacts with the other Fab in a dimer, whereas the corresponding N53 of 3.3-Fab does not. This difference in the protein-protein interaction between two Fab molecules in a dimer may explain the temperature dependence of 2B5 in PEG binding, as well as its inhibition by crown ether.
- Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan. chengung@gate.sinica.edu.tw.
Organizational Affiliation: