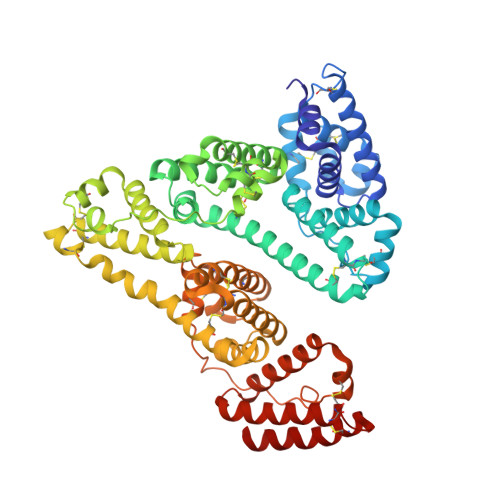
Novel Brain-Tumor-Inhibiting Copper(II) Compound Based on a Human Serum Albumin (HSA)-Cell Penetrating Peptide Conjugate.
Zhang, Z., Yu, P., Gou, Y., Zhang, J., Li, S., Cai, M., Sun, H., Yang, F., Liang, H.(2019) J Med Chem 62: 10630-10644
- PubMed: 31693353 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00939
- Primary Citation Related Structures:
6L4K - PubMed Abstract:
It is a great challenge to design drugs that penetrate the blood-brain barrier to inhibit brain tumor growth by acting against multiple targets and also improve their delivery efficacy and targeting ability to cancer cells. To overcome the above problems, we designed a multitarget metal agent for treating brain tumors based on an human serum albumin (HSA)-cell penetrating peptide conjugate. Thus, we rationally screened copper (Cu) and 2-acetyl-3-ethylpyrazine thiosemicarbazones to synthesize six compounds, and we investigated their structure-activity relationships and confirmed multiple mechanisms for brain glioma cells. The HSA- 6b complex structure indicated that 6b binds to the IIA subdomain of HSA and His242 replaces the Br ligand in 6b in coordination with Cu 2+ . In vivo data suggested that both 6b and the HSA- 6b -peptide conjugate penetrate the blood-brain barrier and inhibit brain tumor growth with few side effects. Furthermore, the HSA-peptide conjugate also improved the delivery efficacy and targeting ability of 6b in vivo.
- State Key Laboratory for the Chemistry and Molecular Engineering of Medicinal Resources , Guangxi Normal University , Guilin , Guangxi 541004 , P. R. China.
Organizational Affiliation: