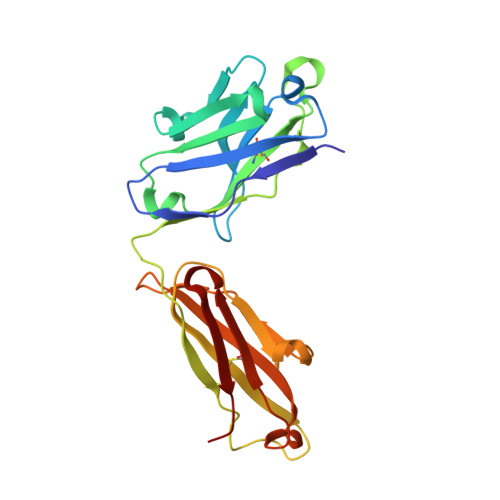
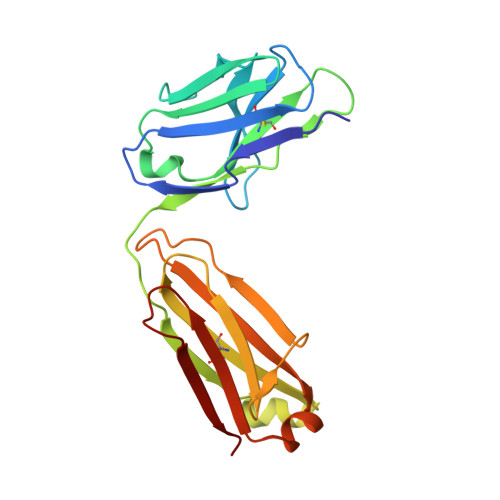
Capturing the inherent structural dynamics of the HIV-1 envelope glycoprotein fusion peptide.
Kumar, S., Sarkar, A., Pugach, P., Sanders, R.W., Moore, J.P., Ward, A.B., Wilson, I.A.(2019) Nat Commun 10: 763-763
- PubMed: 30770829
- DOI: https://doi.org/10.1038/s41467-019-08738-5
- Primary Citation of Related Structures:
6MCO, 6MDT, 6ME1 - PubMed Abstract:
The N-terminal fusion peptide (FP) of the human immunodeficiency virus (HIV)-1 envelope glycoprotein (Env) gp41 subunit plays a critical role in cell entry. However, capturing the structural flexibility in the unbound FP is challenging in the native Env trimer. Here, FP conformational isomerism is observed in two crystal structures of a soluble clade B transmitted/founder virus B41 SOSIP.664 Env with broadly neutralizing antibodies (bNAbs) PGT124 and 35O22 to aid in crystallization and that are not specific for binding to the FP. Large rearrangements in the FP and fusion peptide proximal region occur around M530, which remains anchored in the tryptophan clasp (gp41 W623, W628, W631) in the B41 Env prefusion state. Further, we redesigned the FP at position 518 to reinstate the bNAb VRC34.01 epitope. These findings provide further structural evidence for the dynamic nature of the FP and how a bNAb epitope can be restored during vaccine design.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, 92037, USA.
Organizational Affiliation: