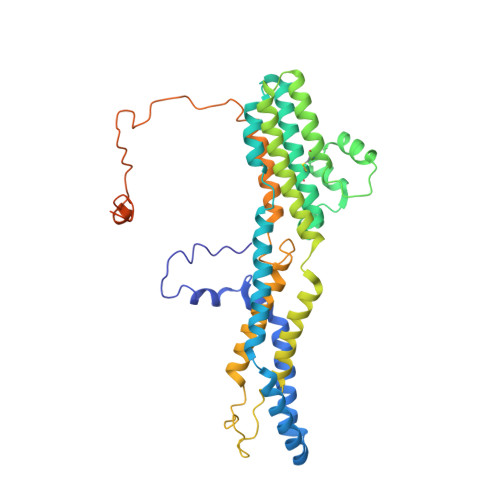
Molecular mechanisms of gating in the calcium-activated chloride channel bestrophin.
Miller, A.N., Vaisey, G., Long, S.B.(2019) Elife 8
- PubMed: 30628889
- DOI: https://doi.org/10.7554/eLife.43231
- Primary Citation Related Structures:
6N23, 6N24, 6N25, 6N26, 6N27, 6N28 - PubMed Abstract:
Bestrophin (BEST1-4) ligand-gated chloride (Cl - ) channels are activated by calcium (Ca 2+ ). Mutation of BEST1 causes retinal disease. Partly because bestrophin channels have no sequence or structural similarity to other ion channels, the molecular mechanisms underlying gating are unknown. Here, we present a series of cryo-electron microscopy structures of chicken BEST1, determined at 3.1 Å resolution or better, that represent the channel's principal gating states. Unlike other channels, opening of the pore is due to the repositioning of tethered pore-lining helices within a surrounding protein shell that dramatically widens a neck of the pore through a concertina of amino acid rearrangements. The neck serves as both the activation and the inactivation gate. Ca 2+ binding instigates opening of the neck through allosteric means whereas inactivation peptide binding induces closing. An aperture within the otherwise wide pore controls anion permeability. The studies define a new molecular paradigm for gating among ligand-gated ion channels.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, United States.
Organizational Affiliation: