Intracellular Neutralization of Ricin Toxin by Single-domain Antibodies Targeting the Active Site.
Rudolph, M.J., Czajka, T.F., Davis, S.A., Thi Nguyen, C.M., Li, X.P., Tumer, N.E., Vance, D.J., Mantis, N.J.(2020) J Mol Biology 432: 1109-1125
- PubMed: 31931008 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2020.01.006
- Primary Citation Related Structures:
6OBC, 6OBE, 6OBG, 6OBM, 6OBO, 6OCA, 6OCD - PubMed Abstract:
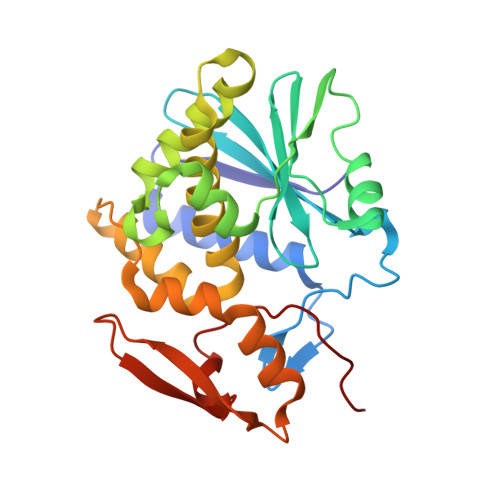
The extreme potency of the plant toxin, ricin, is due to its enzymatic subunit, RTA, which inactivates mammalian ribosomes with near-perfect efficiency. Here we characterized, at the functional and structural levels, seven alpaca single-domain antibodies (V H Hs) previously reported to recognize epitopes in proximity to RTA's active site. Three of the V H Hs, V2A11, V8E6, and V2G10, were potent inhibitors of RTA in vitro and protected Vero cells from ricin when expressed as intracellular antibodies ("intrabodies"). Crystal structure analysis revealed that the complementarity-determining region 3 (CDR3) elements of V2A11 and V8E6 penetrate RTA's active site and interact with key catalytic residues. V2G10, by contrast, sits atop the enzymatic pocket and occludes substrate accessibility. The other four V H Hs also penetrated/occluded RTA's active site, but lacked sufficient binding affinities to outcompete RTA-ribosome interactions. Intracellular delivery of high-affinity, single-domain antibodies may offer a new avenue in the development of countermeasures against ricin toxin.toxin, antibody, structure, intracellular.
- New York Structural Biology Center, New York, NY 10027, United States. Electronic address: mrudolph@nysbc.org.
Organizational Affiliation: