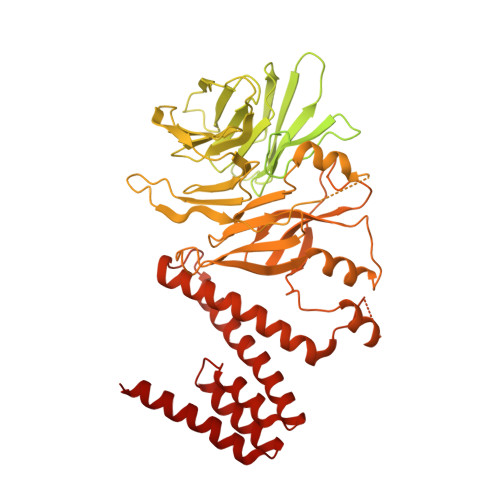
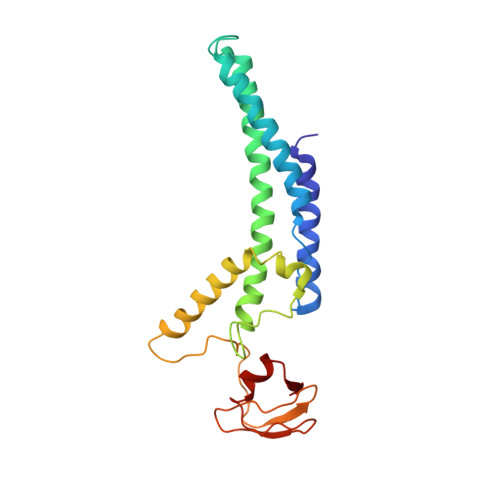
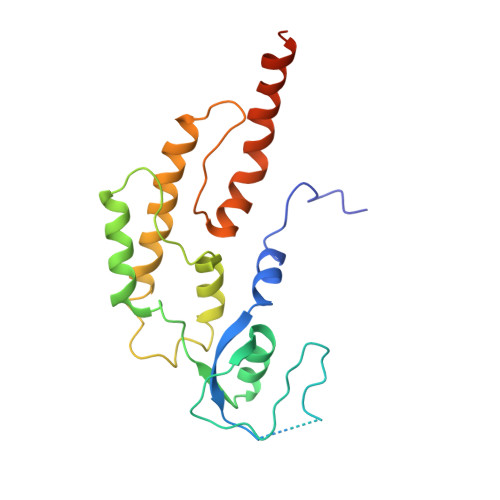
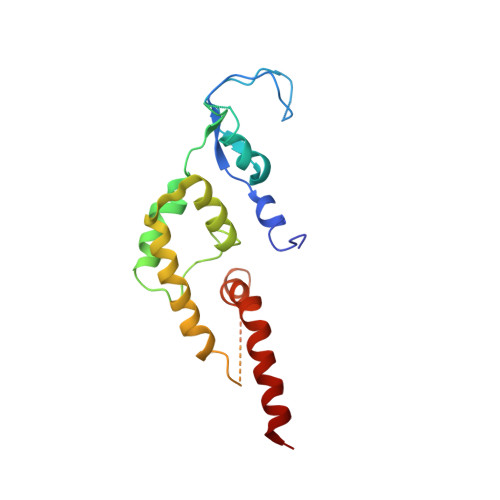
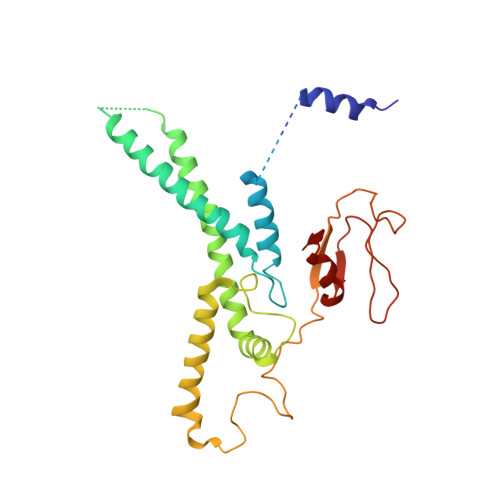
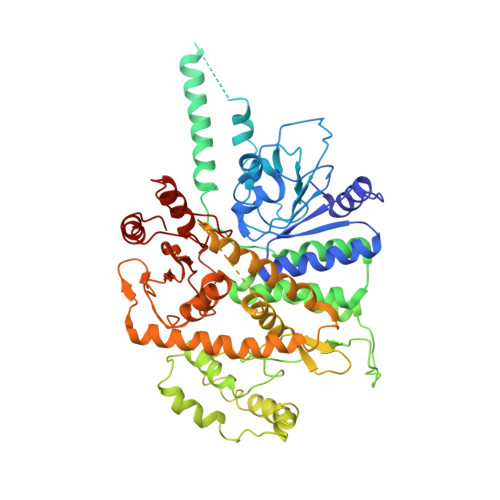
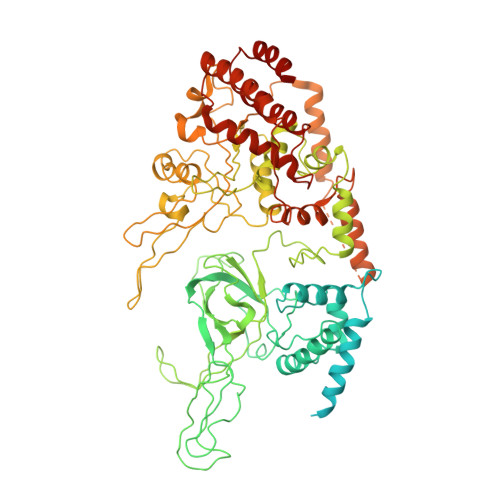
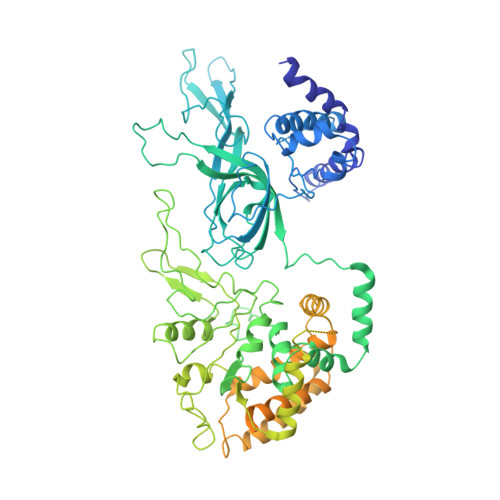
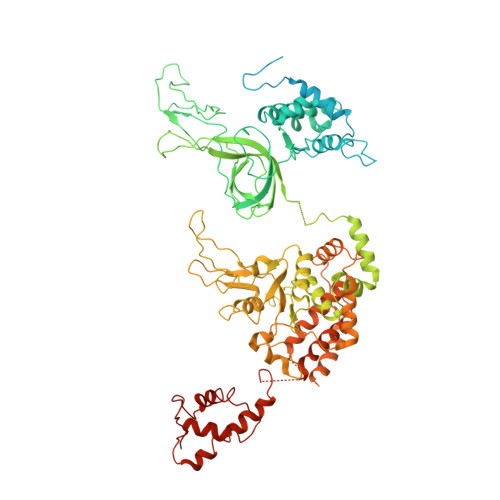
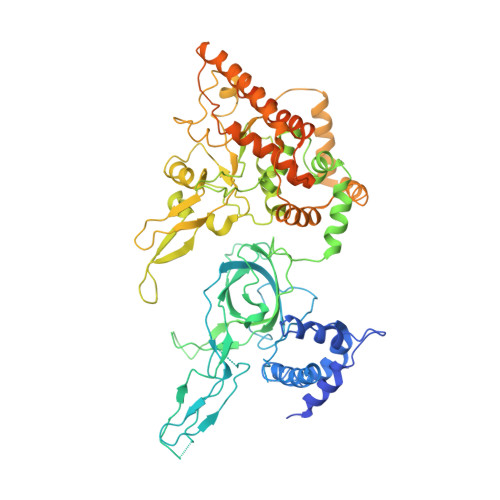
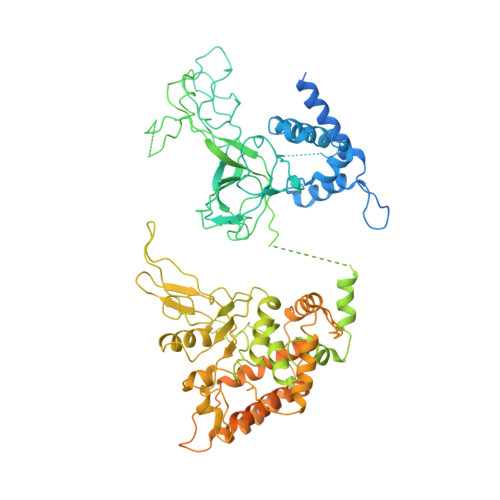
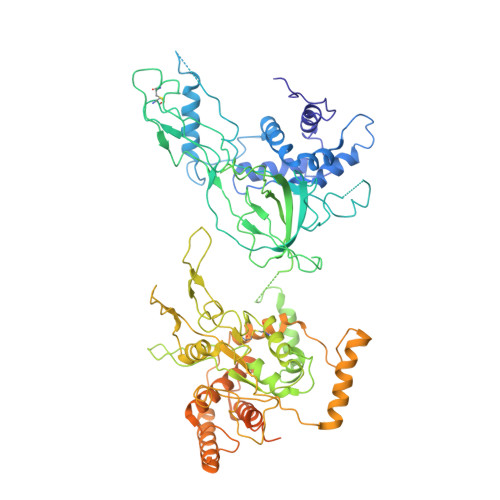
Ctf4 organizes sister replisomes and Pol alpha into a replication factory.
Yuan, Z., Georgescu, R., Santos, R.L.A., Zhang, D., Bai, L., Yao, N.Y., Zhao, G., O'Donnell, M.E., Li, H.(2019) Elife 8
- PubMed: 31589141
- DOI: https://doi.org/10.7554/eLife.47405
- Primary Citation of Related Structures:
6PTJ, 6PTN, 6PTO - PubMed Abstract:
The current view is that eukaryotic replisomes are independent. Here we show that Ctf4 tightly dimerizes CMG helicase, with an extensive interface involving Psf2, Cdc45, and Sld5. Interestingly, Ctf4 binds only one Pol α-primase. Thus, Ctf4 may have evolved as a trimer to organize two helicases and one Pol α-primase into a replication factory. In the 2CMG-Ctf4 3 -1Pol α-primase factory model, the two CMGs nearly face each other, placing the two lagging strands toward the center and two leading strands out the sides. The single Pol α-primase is centrally located and may prime both sister replisomes. The Ctf4-coupled-sister replisome model is consistent with cellular microscopy studies revealing two sister forks of an origin remain attached and are pushed forward from a protein platform. The replication factory model may facilitate parental nucleosome transfer during replication.
- Structural Biology Program, Van Andel Institute, Grand Rapids, United States.
Organizational Affiliation: