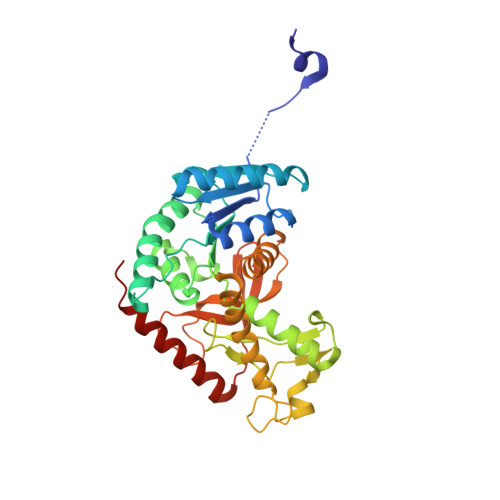
Structural Evidence for Isoform-Selective Allosteric Inhibition of Lactate Dehydrogenase A.
Friberg, A., Rehwinkel, H., Nguyen, D., Putter, V., Quanz, M., Weiske, J., Eberspacher, U., Heisler, I., Langer, G.(2020) ACS Omega 5: 13034-13041
- PubMed: 32548488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.0c00715
- Primary Citation Related Structures:
6SBU, 6SBV - PubMed Abstract:
Lactate dehydrogenase A (LDHA) is frequently overexpressed in tumors, thereby sustaining high glycolysis rates, tumor growth, and chemoresistance. High-throughput screening resulted in the identification of phthalimide and dibenzofuran derivatives as novel lactate dehydrogenase inhibitors, selectively inhibiting the activity of the LDHA isoenzyme. Cocrystallization experiments confirmed target engagement in addition to demonstrating binding to a novel allosteric binding site present in all four LDHA subunits of the LDH5 homotetramer.
- Bayer AG, Pharmaceuticals, R&D, Müllerstrasse 178, 13342 Berlin, Germany.
Organizational Affiliation: