Inhibitor peptides against Mcl-1 containing non-natural amino acids show potent apoptotic response.
Mandal, T., Grant, R.A., Keating, A.E.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Induced myeloid leukemia cell differentiation protein Mcl-1 | 156 | Homo sapiens | Mutation(s): 0 Gene Names: MCL1, BCL2L3 | 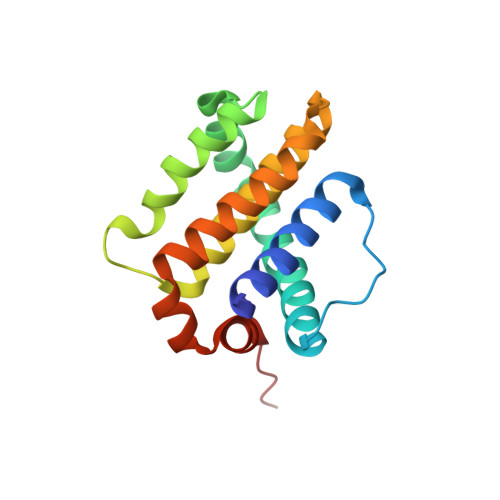 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q07820 GTEx: ENSG00000143384 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q07820 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| modified unnatural Bim BH3 peptide | 23 | Homo sapiens | Mutation(s): 0 | 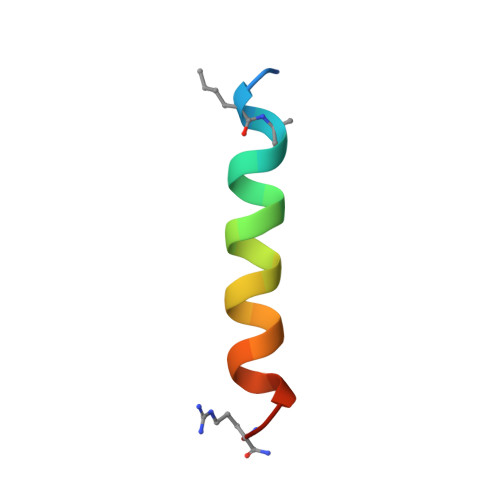 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O43521 GTEx: ENSG00000153094 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O43521 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ADM (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth B] | ADAMANTANE C10 H16 ORILYTVJVMAKLC-YNFQOJQRSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| AIB Query on AIB | B | L-PEPTIDE LINKING | C4 H9 N O2 |  | ALA |
| NLE Query on NLE | B | L-PEPTIDE LINKING | C6 H13 N O2 |  | LEU |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 30.746 | α = 90 |
| b = 56.56 | β = 90 |
| c = 99.515 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Department of Defense (DOD, United States) | United States | -- |