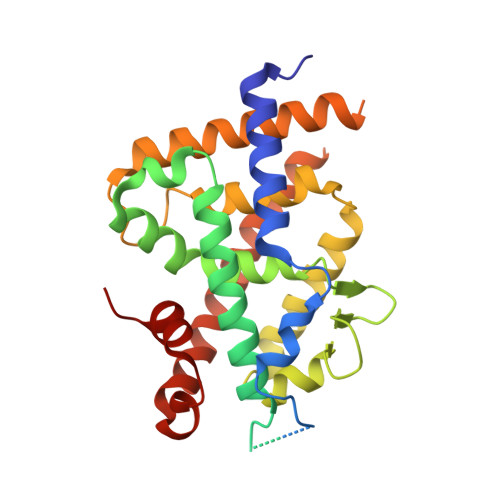
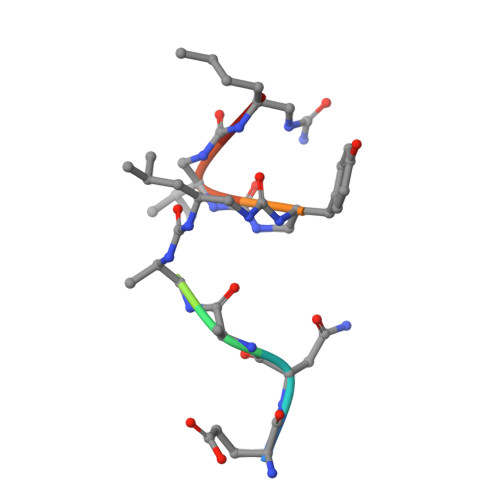
Structural Basis for alpha-Helix Mimicry and Inhibition of Protein-Protein Interactions with Oligourea Foldamers.
Cussol, L., Mauran-Ambrosino, L., Buratto, J., Belorusova, A.Y., Neuville, M., Osz, J., Fribourg, S., Fremaux, J., Dolain, C., Goudreau, S.R., Rochel, N., Guichard, G.(2021) Angew Chem Int Ed Engl 60: 2296-2303
- PubMed: 32935897
- DOI: https://doi.org/10.1002/anie.202008992
- Primary Citation Related Structures:
6HFA, 6XZH, 6XZI, 6XZJ, 6XZK, 6XZV - PubMed Abstract:
Efficient optimization of a peptide lead into a drug candidate frequently needs further transformation to augment properties such as bioavailability. Among the different options, foldamers, which are sequence-based oligomers with precise folded conformation, have emerged as a promising technology. We introduce oligourea foldamers to reduce the peptide character of inhibitors of protein-protein interactions (PPI). However, the precise design of such mimics is currently limited by the lack of structural information on how these foldamers adapt to protein surfaces. We report a collection of X-ray structures of peptide-oligourea hybrids in complex with ubiquitin ligase MDM2 and vitamin D receptor and show how such hybrid oligomers can be designed to bind with high affinity to protein targets. This work should enable the generation of more effective foldamer-based disruptors of PPIs in the context of peptide lead optimization.
- Univ. Bordeaux, CNRS, Bordeaux INP, CBMN, UMR 5248, Institut Européen de Chimie et Biologie, 2 rue Robert Escarpit, F-33607, Pessac, France.
Organizational Affiliation: