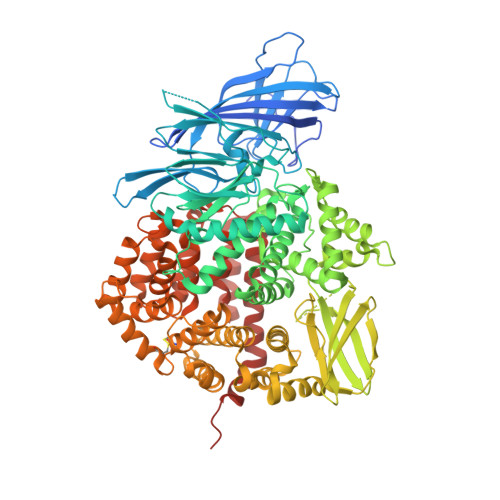
Structural Basis of Inhibition of Insulin-Regulated Aminopeptidase by a Macrocyclic Peptidic Inhibitor.
Mpakali, A., Saridakis, E., Giastas, P., Maben, Z., Stern, L.J., Larhed, M., Hallberg, M., Stratikos, E.(2020) ACS Med Chem Lett 11: 1429-1434
- PubMed: 32676150
- DOI: https://doi.org/10.1021/acsmedchemlett.0c00172
- Primary Citation of Related Structures:
6YDX - PubMed Abstract:
Insulin-regulated aminopeptidase (IRAP) is a transmembrane zinc metallopeptidase with many important biological functions and an emerging pharmacological target. Although previous structural studies have given insight on how IRAP recognizes linear peptides, how it recognizes its physiological cyclic ligands remains elusive. Here, we report the first crystal structure of IRAP with the macrocyclic peptide inhibitor HA08 that combines structural elements from angiotensin IV and the physiological substrates oxytocin and vasopressin. The compound is found in the catalytic site in a near canonical substrate-like configuration and inhibits by a competitive mechanism. Comparison with previously solved structures of IRAP along with small-angle X-ray scattering experiments suggests that IRAP is in an open conformation in solution but undergoes a closing conformational change upon inhibitor binding. Stabilization of the closed conformation in combination with catalytic water exclusion by the tightly juxtaposed GAMEN loop is proposed as a mechanism of inhibition.
- National Center for Scientific Research Demokritos, Agia Paraskevi, Athens 15341, Greece.
Organizational Affiliation: