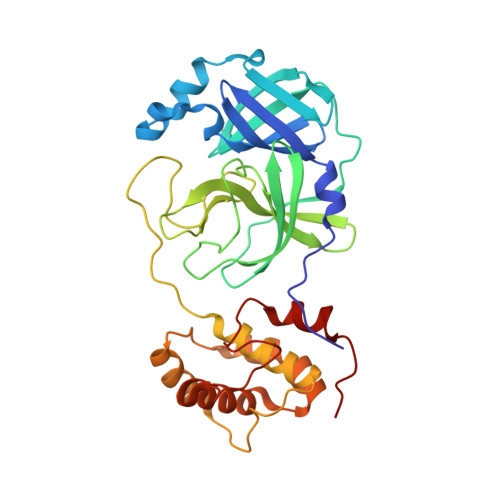
Structural basis for the inhibition of SARS-CoV-2 main protease by antineoplastic drug carmofur.
Jin, Z., Zhao, Y., Sun, Y., Zhang, B., Wang, H., Wu, Y., Zhu, Y., Zhu, C., Hu, T., Du, X., Duan, Y., Yu, J., Yang, X., Yang, X., Yang, K., Liu, X., Guddat, L.W., Xiao, G., Zhang, L., Yang, H., Rao, Z.(2020) Nat Struct Mol Biol 27: 529-532
- PubMed: 32382072
- DOI: https://doi.org/10.1038/s41594-020-0440-6
- Primary Citation Related Structures:
7BUY - PubMed Abstract:
The antineoplastic drug carmofur is shown to inhibit the SARS-CoV-2 main protease (M pro ). Here, the X-ray crystal structure of M pro in complex with carmofur reveals that the carbonyl reactive group of carmofur is covalently bound to catalytic Cys145, whereas its fatty acid tail occupies the hydrophobic S2 subsite. Carmofur inhibits viral replication in cells (EC 50 = 24.30 μM) and is a promising lead compound to develop new antiviral treatment for COVID-19.
- Shanghai Institute for Advanced Immunochemical Studies and School of Life Science and Technology, ShanghaiTech University, Shanghai, China.
Organizational Affiliation: