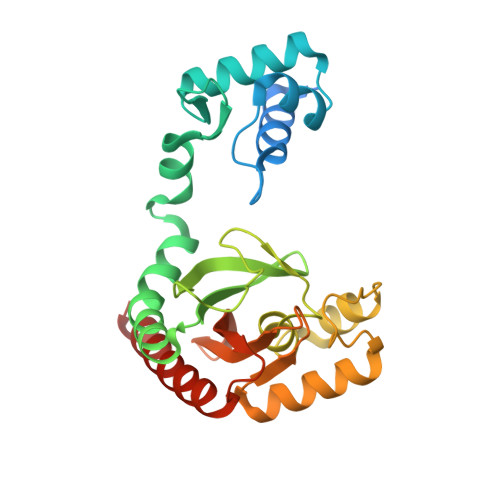
Structural basis of the conformational changes in Microbacterium hydrocarbonoxydans IclR transcription factor homolog due to ligand binding.
Akiyama, T., Sasaki, Y., Ito, S., Yajima, S.(2021) Biochim Biophys Acta Proteins Proteom 1869: 140644-140644
- PubMed: 33716191 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2021.140644
- Primary Citation Related Structures:
7CUO, 7DQB - PubMed Abstract:
Microbacterium hydrocarbonoxydans has been isolated using an unnatural acylhydrazide compound as the sole carbon source. The compound is hydrolyzed by bacterial hydrazidase, and the gene expression of the enzyme is considered to be controlled by a transcription factor of the Isocitrate lyase Regulator (IclR) family, belonging to the one-component signaling systems. Recently, we reported the crystal structure of an unliganded IclR homolog from M. hydrocarbonoxydans, named putative 4-hydroxybenzoate response regulator (pHbrR), which has a unique homotetramer conformation. In this study, we report the crystal structure of pHbrR complexed with 4-hydroxybenzoic acid, the catalytic product of hydrazidase, at 2.0 Å resolution. pHbrR forms a homodimer with multimeric rearrangement in the unliganded state. Gel filtration column chromatography results suggested dimer-tetramer rearrangement. We observed conformational change in the loop region covering the ligand-binding site, and domain rearrangements in the monomer. This study reports the first liganded IclR family protein structure that demonstrates large structural rearrangements between liganded and unliganded proteins, which may represent a general model for IclRs.
- Department of Bioscience, Tokyo University of Agriculture, Setagaya-ku, Tokyo 156-8502, Japan.
Organizational Affiliation: