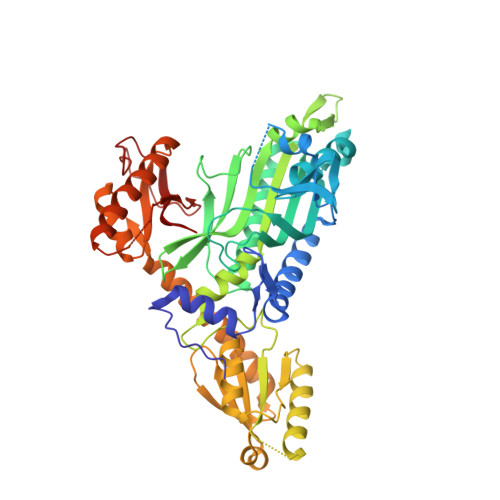
Targeting prolyl-tRNA synthetase via a series of ATP-mimetics to accelerate drug discovery against toxoplasmosis.
Yogavel, M., Bougdour, A., Mishra, S., Malhotra, N., Chhibber-Goel, J., Bellini, V., Harlos, K., Laleu, B., Hakimi, M.A., Sharma, A.(2023) PLoS Pathog 19: e1011124-e1011124
- PubMed: 36854028
- DOI: https://doi.org/10.1371/journal.ppat.1011124
- Primary Citation Related Structures:
7F98, 7F99, 7F9A, 7F9B, 7F9C, 7F9D, 7FAK, 7FAL, 7FAM, 7FAN, 7VC1, 7VC2, 7VC3 - PubMed Abstract:
The prolyl-tRNA synthetase (PRS) is a validated drug target for febrifugine and its synthetic analog halofuginone (HFG) against multiple apicomplexan parasites including Plasmodium falciparum and Toxoplasma gondii. Here, a novel ATP-mimetic centered on 1-(pyridin-4-yl) pyrrolidin-2-one (PPL) scaffold has been validated to bind to Toxoplasma gondii PRS and kill toxoplasma parasites. PPL series exhibited potent inhibition at the cellular (T. gondii parasites) and enzymatic (TgPRS) levels compared to the human counterparts. Cell-based chemical mutagenesis was employed to determine the mechanism of action via a forward genetic screen. Tg-resistant parasites were analyzed with wild-type strain by RNA-seq to identify mutations in the coding sequence conferring drug resistance by computational analysis of variants. DNA sequencing established two mutations, T477A and T592S, proximal to terminals of the PPL scaffold and not directly in the ATP, tRNA, or L-pro sites, as supported by the structural data from high-resolution crystal structures of drug-bound enzyme complexes. These data provide an avenue for structure-based activity enhancement of this chemical series as anti-infectives.
- Molecular Medicine-Structural Parasitology Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), Aruna Asaf Ali Marg, New Delhi, India.
Organizational Affiliation: