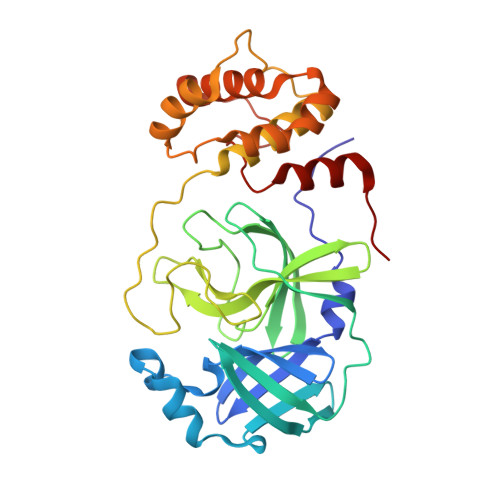
Crystallographic structure of wild-type SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site.
Lee, J., Worrall, L.J., Vuckovic, M., Rosell, F.I., Gentile, F., Ton, A.T., Caveney, N.A., Ban, F., Cherkasov, A., Paetzel, M., Strynadka, N.C.J.(2020) Nat Commun 11: 5877-5877
- PubMed: 33208735
- DOI: https://doi.org/10.1038/s41467-020-19662-4
- Primary Citation Related Structures:
7JOY, 7JP1, 7KHP - PubMed Abstract:
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), the pathogen that causes the disease COVID-19, produces replicase polyproteins 1a and 1ab that contain, respectively, 11 or 16 nonstructural proteins (nsp). Nsp5 is the main protease (M pro ) responsible for cleavage at eleven positions along these polyproteins, including at its own N- and C-terminal boundaries, representing essential processing events for subsequent viral assembly and maturation. We have determined X-ray crystallographic structures of this cysteine protease in its wild-type free active site state at 1.8 Å resolution, in its acyl-enzyme intermediate state with the native C-terminal autocleavage sequence at 1.95 Å resolution and in its product bound state at 2.0 Å resolution by employing an active site mutation (C145A). We characterize the stereochemical features of the acyl-enzyme intermediate including critical hydrogen bonding distances underlying catalysis in the Cys/His dyad and oxyanion hole. We also identify a highly ordered water molecule in a position compatible for a role as the deacylating nucleophile in the catalytic mechanism and characterize the binding groove conformational changes and dimerization interface that occur upon formation of the acyl-enzyme. Collectively, these crystallographic snapshots provide valuable mechanistic and structural insights for future antiviral therapeutic development including revised molecular docking strategies based on M pro inhibition.
- Department of Biochemistry and Molecular Biology and Centre for Blood Research, The University of British Columbia, Vancouver, BC, Canada.
Organizational Affiliation: