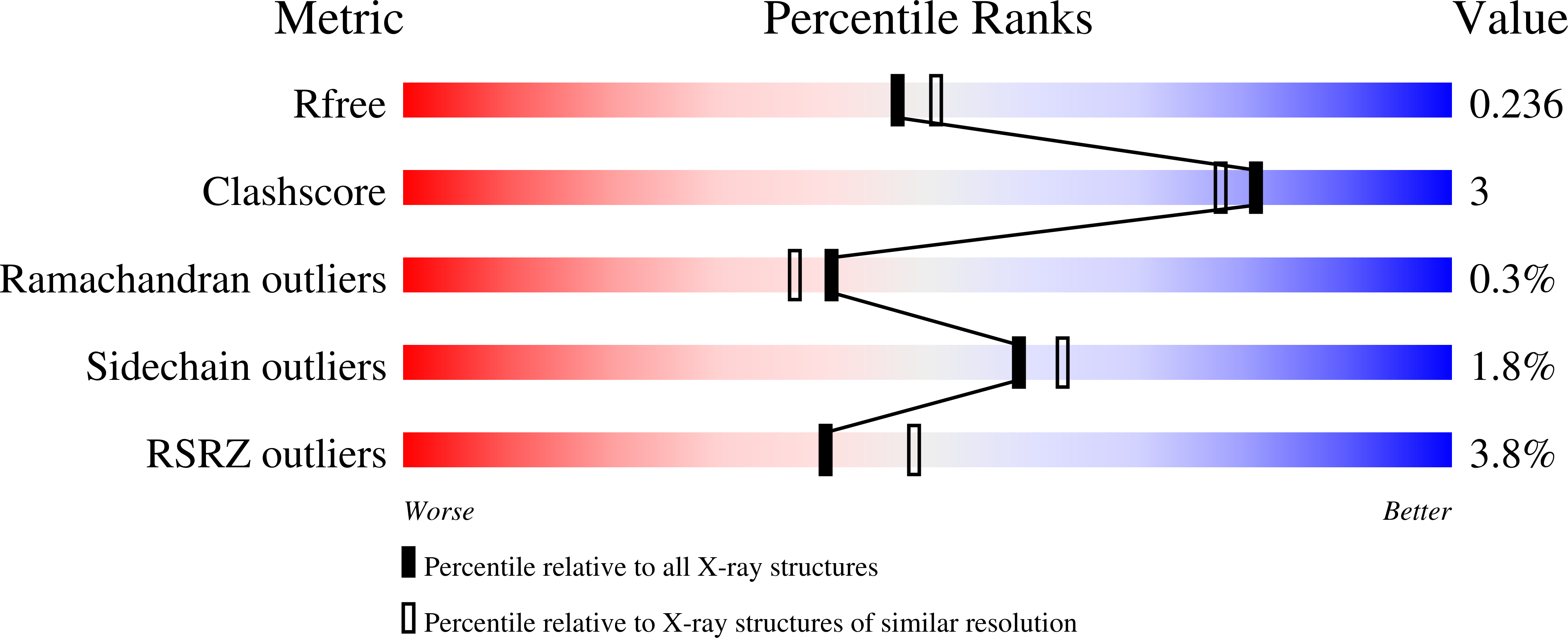
A BCMA/CD16A bispecific innate cell engager for the treatment of multiple myeloma.
Kakiuchi-Kiyota, S., Ross, T., Wallweber, H.A., Kiefer, J.R., Schutten, M.M., Adedeji, A.O., Cai, H., Hendricks, R., Cohen, S., Myneni, S., Liu, L., Fullerton, A., Corr, N., Yu, L., de Almeida Nagata, D., Zhong, S., Leong, S.R., Li, J., Nakamura, R., Sumiyoshi, T., Li, J., Ovacik, A.M., Zheng, B., Dillon, M., Spiess, C., Wingert, S., Rajkovic, E., Ellwanger, K., Reusch, U., Polson, A.G.(2022) Leukemia 36: 1006-1014
- PubMed: 35001074
- DOI: https://doi.org/10.1038/s41375-021-01478-w
- Primary Citation of Related Structures:
7SEG - PubMed Abstract:
Despite the recent progress, multiple myeloma (MM) is still essentially incurable and there is a need for additional effective treatments with good tolerability. RO7297089 is a novel bispecific BCMA/CD16A-directed innate cell engager (ICE ® ) designed to induce BCMA+ MM cell lysis through high affinity binding of CD16A and retargeting of NK cell cytotoxicity and macrophage phagocytosis. Unlike conventional antibodies approved in MM, RO7297089 selectively targets CD16A with no binding of other Fcγ receptors, including CD16B on neutrophils, and irrespective of 158V/F polymorphism, and its activity is less affected by competing IgG suggesting activity in the presence of M-protein. Structural analysis revealed this is due to selective interaction with a single residue (Y140) uniquely present in CD16A opposite the Fc binding site. RO7297089 induced tumor cell killing more potently than conventional antibodies (wild-type and Fc-enhanced) and induced lysis of BCMA+ cells at very low effector-to-target ratios. Preclinical toxicology data suggested a favorable safety profile as in vitro cytokine release was minimal and no RO7297089-related mortalities or adverse events were observed in cynomolgus monkeys. These data suggest good tolerability and the potential of RO7297089 to be a novel effective treatment of MM patients.
Organizational Affiliation:
Genentech Research and Early Development, San Francisco, CA, USA.