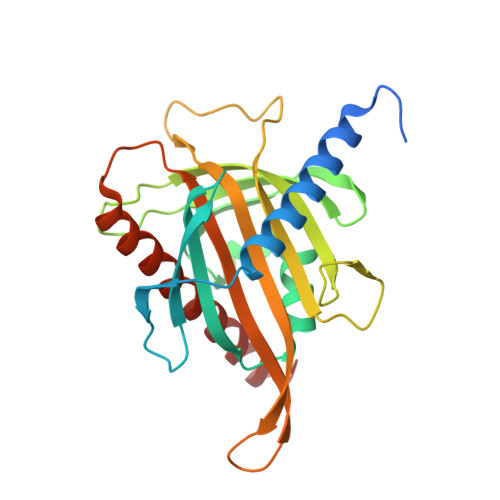
Structural basis for the carotenoid binding and transport function of a START domain.
Sluchanko, N.N., Slonimskiy, Y.B., Egorkin, N.A., Varfolomeeva, L.A., Kleymenov, S.Y., Minyaev, M.E., Faletrov, Y.V., Moysenovich, A.M., Parshina, E.Y., Friedrich, T., Maksimov, E.G., Boyko, K.M., Popov, V.O.(2022) Structure 30: 1647
- PubMed: 36356587
- DOI: https://doi.org/10.1016/j.str.2022.10.007
- Primary Citation of Related Structures:
7ZVQ, 7ZVR - PubMed Abstract:
STARD3, a steroidogenic acute regulatory lipid transfer protein, was identified as a key xanthophyll-binding protein in the human retina. STARD3 and its homologs in invertebrates are known to bind and transport carotenoids, but this lacks structural elucidation. Here, we report high-resolution crystal structures of the apo- and zeaxanthin (ZEA)-bound carotenoid-binding protein from silkworm Bombyx mori (BmCBP). Having a STARD3-like fold, BmCBP features novel elements, including the Ω1-loop that, in the apoform, is uniquely fixed on the α4-helix by an R173-D279 salt bridge. We exploit absorbance, Raman and dichroism spectroscopy, and calorimetry to describe how ZEA and BmCBP mutually affect each other in the complex. We identify key carotenoid-binding residues, confirm their roles by ZEA-binding capacity and X-ray structures of BmCBP mutants, and also demonstrate that markedly different carotenoid-binding capacities of BmCBP and human STARD3 stem from differences in the structural organization of their carotenoid-binding cavity.
Organizational Affiliation:
A.N. Bach Institute of Biochemistry, Federal Research Center of Biotechnology of the Russian Academy of Sciences, 33 Leninsky Prospect, Building 1, Moscow 119071, Russian Federation. Electronic address: nikolai.sluchanko@mail.ru.