Expansion of the Catalytic Repertoire of Alcohol Dehydrogenases in Plant Metabolism.
Langley, C., Tatsis, E., Hong, B., Nakamura, Y., Paetz, C., Stevenson, C.E.M., Basquin, J., Lawson, D.M., Caputi, L., O'Connor, S.E.(2022) Angew Chem Int Ed Engl 61: e202210934-e202210934
- PubMed: 36198083 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202210934
- Primary Citation Related Structures:
8A3N, 8B1V, 8B25, 8B26, 8B27 - PubMed Abstract:
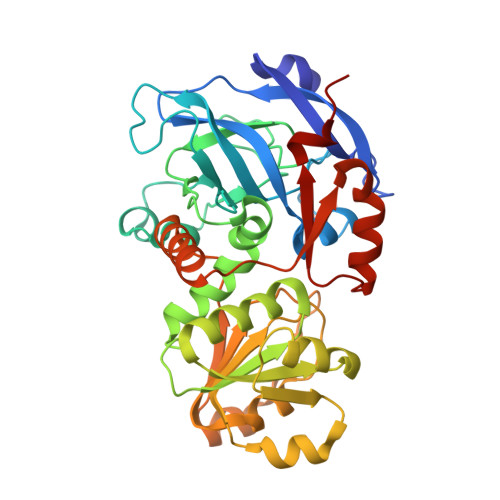
Medium-chain alcohol dehydrogenases (ADHs) comprise a highly conserved enzyme family that catalyse the reversible reduction of aldehydes. However, recent discoveries in plant natural product biosynthesis suggest that the catalytic repertoire of ADHs has been expanded. Here we report the crystal structure of dihydroprecondylocarpine acetate synthase (DPAS), an ADH that catalyses the non-canonical 1,4-reduction of an α,β-unsaturated iminium moiety. Comparison with structures of plant-derived ADHs suggest the 1,4-iminium reduction does not require a proton relay or the presence of a catalytic zinc ion in contrast to canonical 1,2-aldehyde reducing ADHs that require the catalytic zinc and a proton relay. Furthermore, ADHs that catalysed 1,2-iminium reduction required the presence of the catalytic zinc and the loss of the proton relay. This suggests how the ADH active site can be modified to perform atypical carbonyl reductions, providing insight into how chemical reactions are diversified in plant metabolism.
- Department of Natural Product Biosynthesis, Max Planck Institute for Chemical Ecology, Hans-Knöll Straße 8, Jena, 07745, Germany.
Organizational Affiliation: