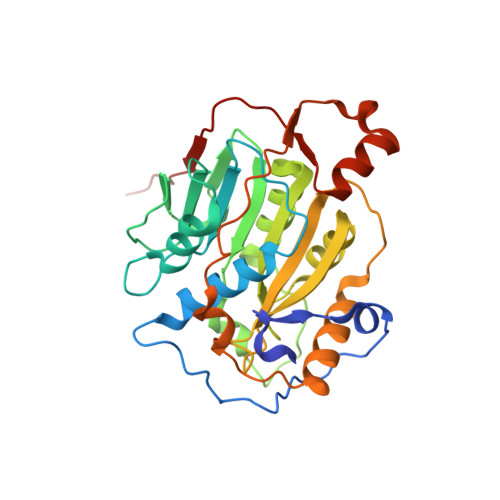
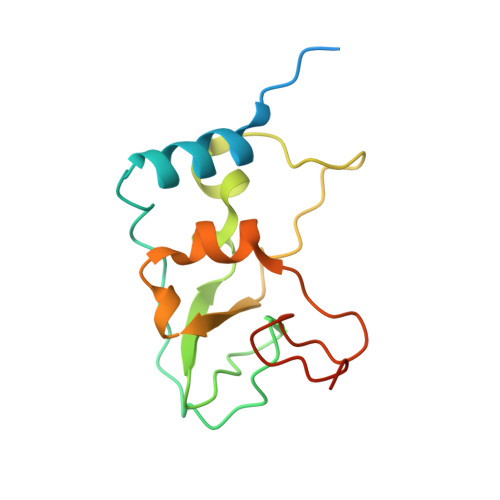
Structural Basis for Inhibition of the SARS-CoV-2 nsp16 by Substrate-Based Dual Site Inhibitors.
Kalnins, G., Rudusa, L., Bula, A.L., Zelencova-Gopejenko, D., Bobileva, O., Sisovs, M., Tars, K., Jirgensons, A., Jaudzems, K., Bobrovs, R.(2024) ChemMedChem 19: e202400618-e202400618
- PubMed: 39258386
- DOI: https://doi.org/10.1002/cmdc.202400618
- Primary Citation of Related Structures:
8RV4, 8RV5, 8RV6, 8RV7, 8RV8, 8RV9, 8RVA, 8RVB, 8RZC, 8RZD, 8RZE - PubMed Abstract:
Coronaviruses, including SARS-CoV-2, possess an mRNA 5' capping apparatus capable of mimicking the natural eukaryotic capping signature. Two SAM-dependent methylating enzymes play important roles in this process: nsp14 methylates the N7 of the guanosine cap, and nsp16-nsp10 methylates the 2'-O- of subsequent nucleotides of viral mRNA. The 2'-O-methylation performed by nsp16-nsp10 is crucial for the escape of the viral RNA from innate immunity. Inhibition of this enzymatic activity has been proposed as a way to combat coronaviruses. In this study, we employed X-ray crystallography to analyze the binding of the SAM analogues to the active site of nsp16-nsp10. We obtained eleven 3D crystal structures of the nsp16-nsp10 complexes with SAM-derived inhibitors, demonstrated different conformations of the methionine substituting part of the molecules, and confirmed that simultaneous dual-site targeting of both SAM and RNA sites correlates with higher inhibitory potential.
- Latvian Biomedical Research and Study Centre, Structural Biology Group, LATVIA.
Organizational Affiliation: