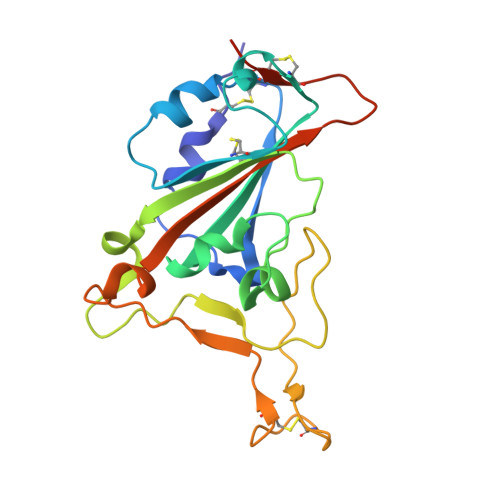
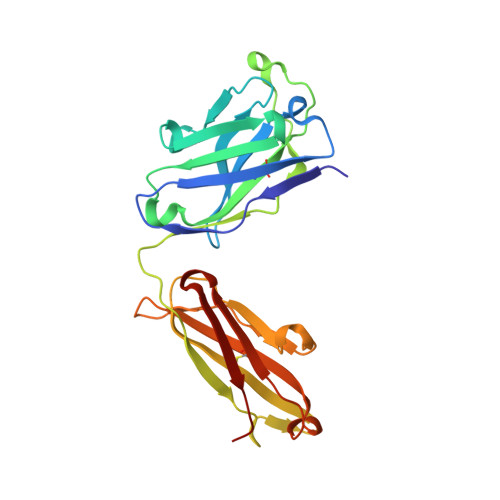
Broadly neutralizing antibodies targeting a conserved silent face of spike RBD resist extreme SARS-CoV-2 antigenic drift
Song, G., Yuan, M., Liu, H., Capozzola, T., Lin, R.N., Torres, J.L., He, W.T., Musharrafieh, R., Dueker, K., Zhou, P., Callaghan, S., Mishra, N., Yong, P., Anzanello, F., Avillion, G., Vo, A.L., Li, X., Zhang, Y., Makhdoomi, M., Feng, Z., Zhu, X., Peng, L., Nemazee, D., Safonova, Y., Briney, B., Ward, A.B., Burton, D.R., Wilson, I.A., Andrabi, R.(2025) Cell Rep 44: 115948