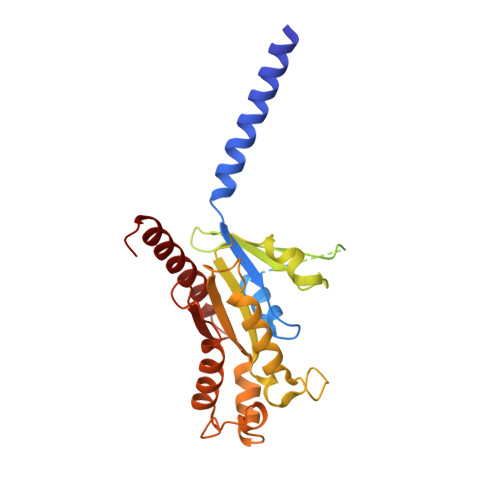
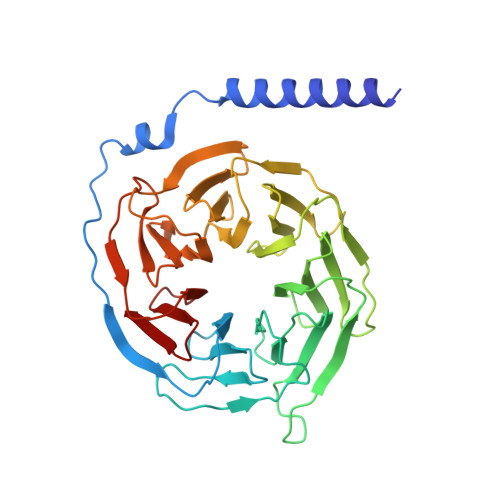
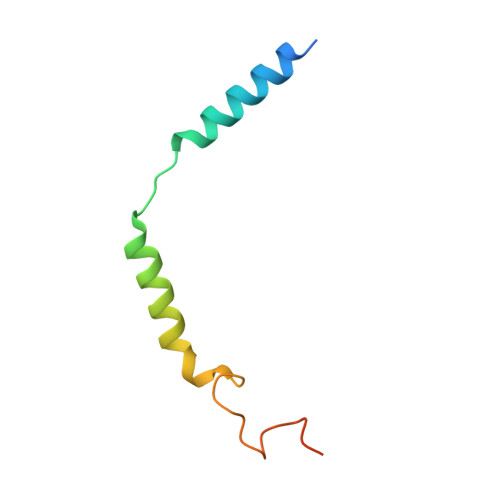
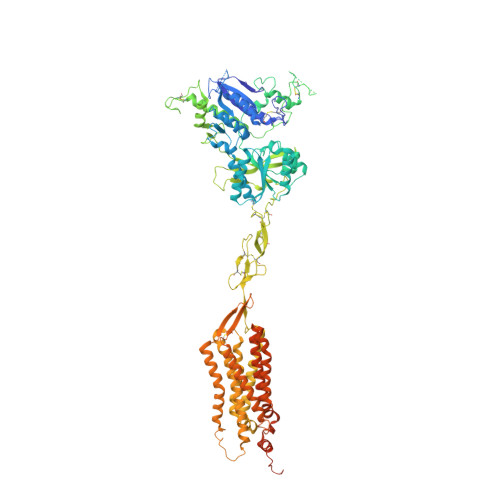
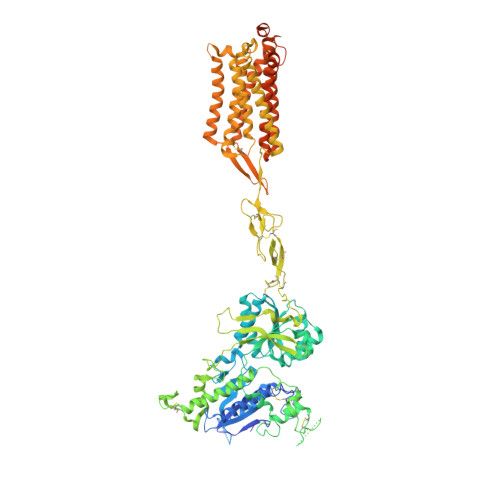
Allosteric modulation and G-protein selectivity of the Ca 2+ -sensing receptor.
He, F., Wu, C.G., Gao, Y., Rahman, S.N., Zaoralova, M., Papasergi-Scott, M.M., Gu, T.J., Robertson, M.J., Seven, A.B., Li, L., Mathiesen, J.M., Skiniotis, G.(2024) Nature 626: 1141-1148
- PubMed: 38326620 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07055-2
- Primary Citation Related Structures:
8SZF, 8SZG, 8SZH, 8SZI - PubMed Abstract:
The calcium-sensing receptor (CaSR) is a family C G-protein-coupled receptor 1 (GPCR) that has a central role in regulating systemic calcium homeostasis 2,3 . Here we use cryo-electron microscopy and functional assays to investigate the activation of human CaSR embedded in lipid nanodiscs and its coupling to functional G i versus G q proteins in the presence and absence of the calcimimetic drug cinacalcet. High-resolution structures show that both G i and G q drive additional conformational changes in the activated CaSR dimer to stabilize a more extensive asymmetric interface of the seven-transmembrane domain (7TM) that involves key protein-lipid interactions. Selective G i and G q coupling by the receptor is achieved through substantial rearrangements of intracellular loop 2 and the C terminus, which contribute differentially towards the binding of the two G-protein subtypes, resulting in distinct CaSR-G-protein interfaces. The structures also reveal that natural polyamines target multiple sites on CaSR to enhance receptor activation by zipping negatively charged regions between two protomers. Furthermore, we find that the amino acid L-tryptophan, a well-known ligand of CaSR extracellular domains, occupies the 7TM bundle of the G-protein-coupled protomer at the same location as cinacalcet and other allosteric modulators. Together, these results provide a framework for G-protein activation and selectivity by CaSR, as well as its allosteric modulation by endogenous and exogenous ligands.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA, USA.
Organizational Affiliation: