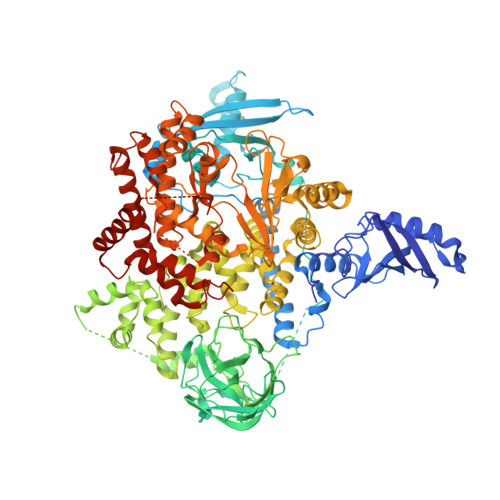
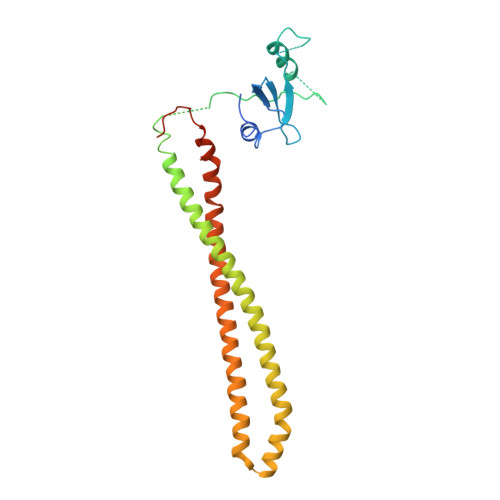
Discovery of Pyridopyrimidinones that Selectively Inhibit the H1047R PI3K alpha Mutant Protein.
Ketcham, J.M., Harwood, S.J., Aranda, R., Aloiau, A.N., Bobek, B.M., Briere, D.M., Burns, A.C., Caddell Haatveit, K., Calinisan, A., Clarine, J., Elliott, A., Engstrom, L.D., Gunn, R.J., Ivetac, A., Jones, B., Kuehler, J., Lawson, J.D., Nguyen, N., Parker, C., Pearson, K.E., Rahbaek, L., Saechao, B., Wang, X., Waters, A., Waters, L., Watkins, A.H., Olson, P., Smith, C.R., Christensen, J.G., Marx, M.A.(2024) J Med Chem 67: 4936-4949
- PubMed: 38477582
- DOI: https://doi.org/10.1021/acs.jmedchem.4c00078
- Primary Citation of Related Structures:
8V8H, 8V8I, 8V8J, 8V8U, 8V8V - PubMed Abstract:
The H1047R mutation of PIK3CA is highly prevalent in breast cancers and other solid tumors. Selectively targeting PI3Kα H1047R over PI3Kα WT is crucial due to the role that PI3Kα WT plays in normal cellular processes, including glucose homeostasis. Currently, only one PI3Kα H1047R -selective inhibitor has progressed into clinical trials, while three pan mutant (H1047R, H1047L, H1047Y, E542K, and E545K) selective PI3Kα inhibitors have also reached the clinical stage. Herein, we report the design and discovery of a series of pyridopyrimidinones that inhibit PI3Kα H1047R with high selectivity over PI3Kα WT , resulting in the discovery of compound 17 . When dosed in the HCC1954 tumor model in mice, 17 provided tumor regressions and a clear pharmacodynamic response. X-ray cocrystal structures from several PI3Kα inhibitors were obtained, revealing three distinct binding modes within PI3Kα H1047R including a previously reported cryptic pocket in the C-terminus of the kinase domain wherein we observe a ligand-induced interaction with Arg1047.
- Mirati Therapeutics, 3545 Cray Court, San Diego, California 92121, United States.
Organizational Affiliation: