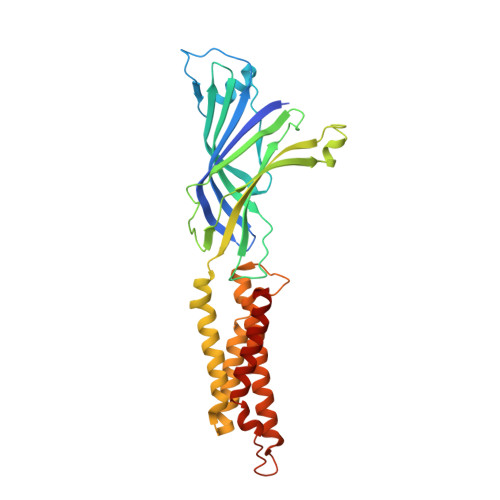
Open-channel structure of a pentameric ligand-gated ion channel reveals a mechanism of leaflet-specific phospholipid modulation.
Petroff, J.T., Dietzen, N.M., Santiago-McRae, E., Deng, B., Washington, M.S., Chen, L.J., Trent Moreland, K., Deng, Z., Rau, M., Fitzpatrick, J.A.J., Yuan, P., Joseph, T.T., Henin, J., Brannigan, G., Cheng, W.W.L.(2022) Nat Commun 13: 7017
- PubMed: 36385237
- DOI: https://doi.org/10.1038/s41467-022-34813-5
- Primary Citation Related Structures:
8D63, 8D64, 8D65, 8D66, 8D67, 8VUW - PubMed Abstract:
Pentameric ligand-gated ion channels (pLGICs) mediate synaptic transmission and are sensitive to their lipid environment. The mechanism of phospholipid modulation of any pLGIC is not well understood. We demonstrate that the model pLGIC, ELIC (Erwinia ligand-gated ion channel), is positively modulated by the anionic phospholipid, phosphatidylglycerol, from the outer leaflet of the membrane. To explore the mechanism of phosphatidylglycerol modulation, we determine a structure of ELIC in an open-channel conformation. The structure shows a bound phospholipid in an outer leaflet site, and structural changes in the phospholipid binding site unique to the open-channel. In combination with streamlined alchemical free energy perturbation calculations and functional measurements in asymmetric liposomes, the data support a mechanism by which an anionic phospholipid stabilizes the activated, open-channel state of a pLGIC by specific, state-dependent binding to this site.
- Department of Anesthesiology, Washington University School of Medicine, Saint Louis, MO, USA.
Organizational Affiliation: