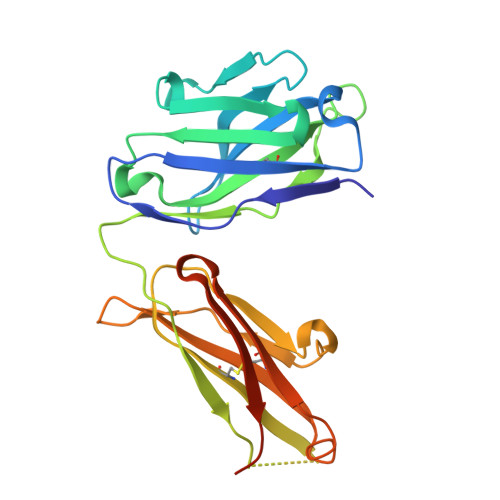
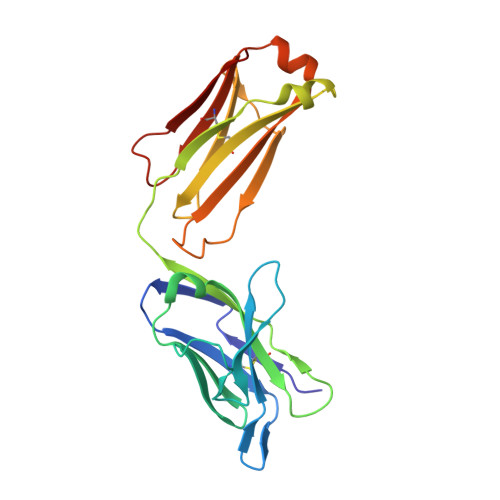
Humanization and engineering of protective antibodies targeting severe fever with thrombocytopenia syndrome virus Gn protein.
Ren, X., Deng, W., Yu, F., Kuang, W., Na, H., Yang, S., Sun, J., Xu, J., Shu, B., Cao, S., Ke, X., Deng, Z., Ning, Y.J., Zhao, H.(2026) Cell Rep 45: 116936-116936
- PubMed: 41671090
- DOI: https://doi.org/10.1016/j.celrep.2026.116936
- Primary Citation of Related Structures:
9VDY, 9VDZ, 9VE0 - PubMed Abstract:
Severe fever with thrombocytopenia syndrome virus (SFTSV) is a highly lethal tick-borne bunyavirus with no approved therapies. We previously developed a murine-human chimeric antibody S2A5, which provides complete protection against lethal SFTSV infection, but its clinical use is limited by potential immunogenicity and moderate activity against certain genotypes. Here, we systematically humanize and optimize S2A5 using five complementary computational platforms, generating eleven variants and identifying hA5-6, which retains potent neutralizing activity and confers full protection in mice. Structure-guided engineering and in silico mutational analysis further improve antibody function, yielding two optimized hA5-6 variants with up to a 317-fold increase in neutralization potency. Biochemical and functional assays, together with cryo-EM reconstruction of an optimized variant bound to SFTSV virions, indicate that the enhanced activity is associated with improved binding to recombinant Gn and intact virions. This study identifies promising SFTSV therapeutics and establishes a generalizable antibody optimization framework.
- State Key Laboratory of Virology and Biosafety, College of Life Sciences, Wuhan University, Wuhan, Hubei, China.
Organizational Affiliation: