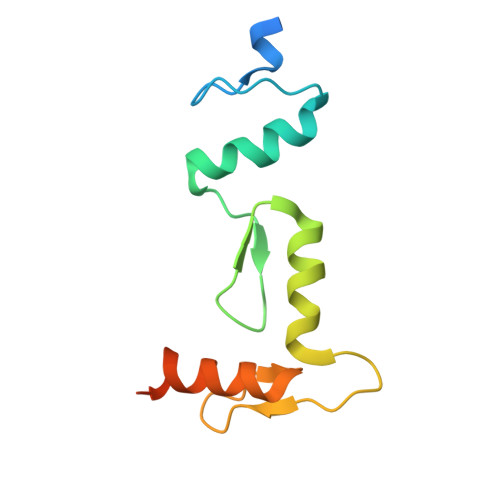
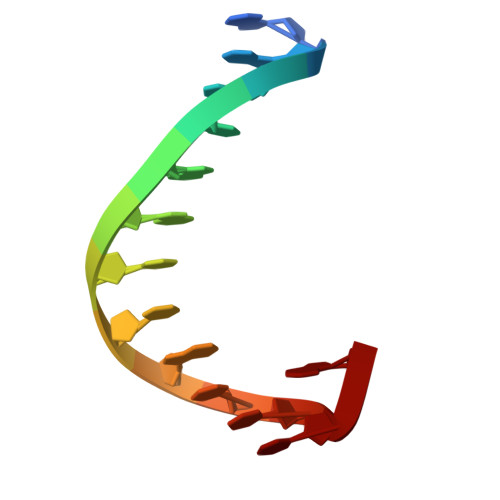
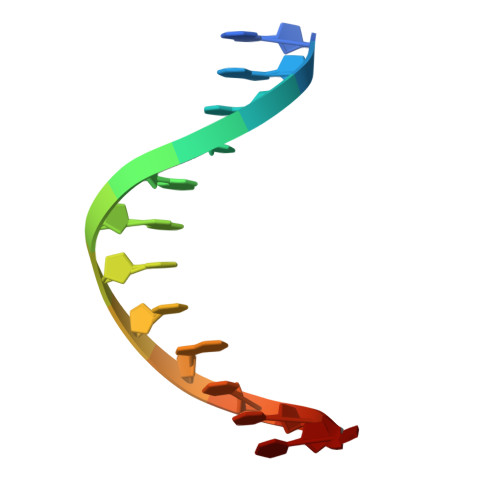
Regulation of BCL11A DNA binding and expression in human erythrocyte precursor HUDEP-2 cells.
Yu, M., Das, P., Horton, J.R., Zhou, J., Lee, J., Hong, T., Lu, Y., Estecio, M.R., Iakova, P.A., Jain, A.K., Sbardella, G., Xiong, Y., Jin, J., Blumenthal, R.M., Huang, Y., Zhang, X., Cheng, X.(2026) bioRxiv
- PubMed: 41822826
- DOI: https://doi.org/10.64898/2026.02.06.704516
- Primary Citation Related Structures:
9YLL, 9YLM, 9YLN, 9YLO, 9YLP - PubMed Abstract:
BCL11A is a transcription factor crucial for neurodevelopment and hematopoiesis. It regulates the developmental switch from fetal hemoglobin (HbF) to adult hemoglobin and is a major therapeutic target for sickle cell disease and β-thalassemia. BCL11A exists in multiple isoforms, including the L isoform (containing a single two-finger ZF2-3 DNA-binding domain) and the XL isoform (containing two arrays: the two-finger ZF2-3 and the three-finger ZF4-6). We used three approaches to investigate BCL11A functions. First, we examined DNA recognition by BCL11A, which preferentially binds the short 6-bp DNA motif TGnCCA. ZF4-6 recognizes all four variants of this motif with distinct strand-specific interactions: TGtCCA on the top strand, TG(A/C)CCA on the complementary strand, and the palindromic TGgCCA on either strand. ZF2-3 also binds TGtCCA from the top strand, featuring a unique thymine interaction by ZF2 Phe388. Motif multiplicity within BCL11A binding sites may promote BCL11A oligomerization by enabling multiple ZF arrays to engage DNA simultaneously. Second, we treated HUDEP-2 cells (which express adult hemoglobin) with inhibitors targeting three epigenetic silencing marks - DNA methylation, histone H3 lysine 9 methylation or H3 lysine 27 methylation. All treatments, individually or in combination, increased HbF expression to varying degrees. Notably, FTX6058 markedly reduced BCL11A transcription and translation (likely via effects on LIN28B), while EML741 caused a partial reduction. Third, we screened 213 pomalidomide- and lenalidomide-derived compounds and quantified proportions of HbF+ cells by flow cytometry. Effects of four compounds were analyzed by protein mass spectrometry. Although BCL11A levels themselves were unchanged, all four compounds selectively decreased levels of known pomalidomide targets, with consistently decreased levels of the zinc-finger proteins IKZF1 and ZFP91. Together, our studies clarify how BCL11A recognizes DNA, how its expression can be modulated epigenetically, and how small-molecule degraders influence its regulatory network, providing new avenues for HbF reactivation therapies.
- Department of Epigenetics and Molecular Carcinogenesis, University of Texas MD Anderson Cancer Center, Houston, TX 77030, USA.
Organizational Affiliation: